- CBSE Class 10 Study Material

CBSE Class 10 Maths Case Study Questions for Maths Chapter 6 (Published by CBSE)
Check case study questions released by cbse for class 10 maths chapter 6 - triangles. solve these questions to prepare the case study questions for the cbse class 10 maths exam 2021-22..

CBSE Class 10 Maths Case Study Questions for Chapter 6 - Triangles are available here. Students must practice with these questions to perform well in their Maths exam. All these case study questions have been published by the Central Board of Secondary Education (CBSE). For the convenience of students, all the questions are provided with answers.
Case Study Questions for Class 10 Maths Chapter 6 - Triangles
CASE STUDY 1:
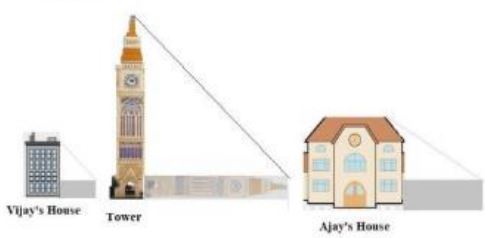
Vijay is trying to find the average height of a tower near his house. He is using the properties of similar triangles.The height of Vijay’s house if 20m when Vijay’s house casts a shadow 10m long on the ground. At the same time, the tower casts a shadow 50m long on the ground and the house of Ajay casts 20m shadow on the ground.
1. What is the height of the tower?
Answer: c) 100m
2. What will be the length of the shadow of the tower when Vijay’s house casts a shadow of 12m?
Answer: d) 60m
3. What is the height of Ajay’s house?
Answer: b) 40m
4. When the tower casts a shadow of 40m, same time what will be the length of the shadow of Ajay’s house?
Answer: a) 16m
5. When the tower casts a shadow of 40m, same time what will be the length of the shadow of Vijay’s house?
Answer: d) 8m
CASE STUDY 2:
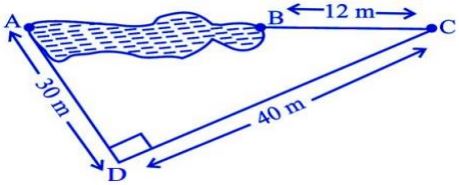
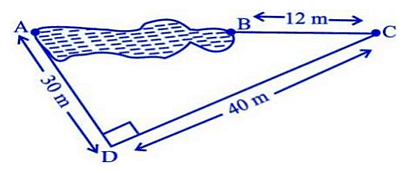
Rohan wants to measure the distance of a pond during the visit to his native. He marks points A and B on the opposite edges of a pond as shown in the figure below. To find the distance between the points, he makes a right-angled triangle using rope connecting B with another point C are a distance of 12m, connecting C to point D at a distance of 40m from point C and the connecting D to the point A which is are a distance of 30m from D such the ∠ ADC=90 0 .
1. Which property of geometry will be used to find the distance AC?
a) Similarity of triangles
b) Thales Theorem
c) Pythagoras Theorem
d) Area of similar triangles
Answer: c)Pythagoras Theorem
2. What is the distance AC?
Answer: a) 50m
3. Which is the following does not form a Pythagoras triplet?
a) (7, 24, 25)
b) (15, 8, 17)
c) (5, 12, 13)
d) (21, 20, 28)
Answer: d) (21, 20, 28)
4. Find the length AB?
Answer: b) 38m
5. Find the length of the rope used.
Answer: c)82m
SCALE FACTOR
Case study:
A scale drawing of an object is the same shape at the object but a different size. The scale of a drawing is a comparison of the length used on a drawing to the length it represents. The scale is written as a ratio. The ratio of two corresponding sides in similar figures is called the scale factor
Scale factor= length in image / corresponding length in object
If one shape can become another using revising, then the shapes are similar. Hence, two shapes are similar when one can become the other after a resize, flip, slide or turn. In the photograph below showing the side view of a train engine. Scale factor is 1:200

This means that a length of 1 cm on the photograph above corresponds to a length of 200cm or 2 m, of the actual engine. The scale can also be written as the ratio of two lengths.
1. If the length of the model is 11cm, then the overall length of the engine in the photograph above, including the couplings (mechanism used to connect) is:
Answer: a)22m
2. What will affect the similarity of any two polygons?
a) They are flipped horizontally
b) They are dilated by a scale factor
c) They are translated down
d) They are not the mirror image of one another.
Answer: d)They are not the mirror image of one another
3. What is the actual width of the door if the width of the door in photograph is 0.35cm?
Answer: a)0.7m
4. If two similar triangles have a scale factor 5:3 which statement regarding the two triangles is true?
a) The ratio of their perimeters is 15:1
b) Their altitudes have a ratio 25:15
c) Their medians have a ratio 10:4
d) Their angle bisectors have a ratio 11:5
Answer: b)Their altitudes have a ratio 25:15
5. The length of AB in the given figure:
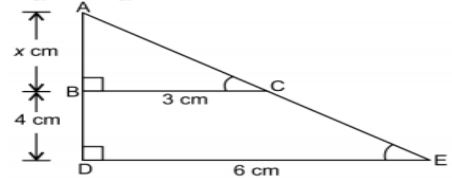
Answer: c)4cm
Also Check:
CBSE Case Study Questions for Class 10 Maths - All Chapters
Tips to Solve Case Study Based Questions Accurately
Get here latest School , CBSE and Govt Jobs notification in English and Hindi for Sarkari Naukari and Sarkari Result . Download the Jagran Josh Sarkari Naukri App .
- Kerala SET Admit Card 2024
- WBCS Prelims Result 2024
- India Post GDS Apply Online 2024
- CTET Answer Key 2024
- Bihar BEd Result 2024
- India Post GDS Vacancy 2024
- Rajasthan BSTC Cut Off 2024
- Bihar STET Answer Key 2024
- SSC CHSL Answer Key 2024
- India Post GDS Cut Off
- Education News
- CBSE Study Material
- CBSE Class 10
Latest Education News
WBBSE West Bengal Board Class 11 Computer Science Syllabus 2024-25: Free PDF Download
NEET UG Counselling 2024 to Begin on July 24, Results to be Announced Tomorrow
Assam Board Class 9 General Maths Syllabus 2024-25: Download Detailed Syllabus PDF For Free!
Today Current Affairs Hindi One Liners: 18 जुलाई 2024- 'कीर्ति' पहल
HSSC Accountant Recruitment 2024: Apply Online For 1296 Various posts, Check Eligibility
भारतीय रेलवे के बारे में कितना जानते हैं आप, पढ़ें क्विज
SSC CHSL 2024 Answer Key: जारी होने वाली है एसएससी टियर 1 की उत्तर कुंजी, यहाँ से करें Download PDF
India Post GDS Recruitment Notification 2024 OUT at indiapostgdsonline.gov.in, 44228 Vacancies Notified, Check Last Date
IAF Agniveer Vayu admit card 2024 for Intake 01/2025 out at agnipathvayu.cdac.in: Check Hall Ticket Download Link
Vocabulary for Bank Exams 2024: 100+ Antonyms and Synonyms for Bank Clerk and PO Exams
IND Vs SL Squad 2024: हार्दिक या सूर्या किसे मिलेगी टीम की कमान? यहां देखें लेटेस्ट टीम अपडेट
HSSC CET Group C Recruitment 2024: हरियाणा में निकली ग्रुप सी के 1838 पदों पर भर्ती, जानें कैसे करें Apply
ICAI CA Final Exam Dates Out at icai.org, Registration Begins on August 7
CSIR NET 2024 Exam Centre: State-wise Test City List and Code
Dr MGR Medical University Result 2024 OUT at tnmgrmu.ac.in; Direct Link to Download UG and PG Marksheet
CSIR NET Previous Year Question Paper, Download PDF with Solutions
PU Result 2024 OUT at puchd.ac.in; Download Panjab University UG and PG Marksheet
UKPSC APS Notification 2024 Out for 99 Vacancies: Apply Online at pcs.uk.gov.in, Check Eligibility Criteria, Application Fees
Today Current Affairs Quiz In Hindi: 18 जुलाई 2024- 'कीर्ति' पहल
NIPER JEE Counselling 2024: MS, M Pharm Seat Matrix Released at niperguwahati.ac.in
CBSE Expert
CBSE Class 10 Maths Case Study Questions PDF
Download Case Study Questions for Class 10 Mathematics to prepare for the upcoming CBSE Class 10 Final Exam. These Case Study and Passage Based questions are published by the experts of CBSE Experts for the students of CBSE Class 10 so that they can score 100% on Boards.

CBSE Class 10 Mathematics Exam 2024 will have a set of questions based on case studies in the form of MCQs. The CBSE Class 10 Mathematics Question Bank on Case Studies, provided in this article, can be very helpful to understand the new format of questions. Share this link with your friends.
Table of Contents
Chapterwise Case Study Questions for Class 10 Mathematics
Inboard exams, students will find the questions based on assertion and reasoning. Also, there will be a few questions based on case studies. In that, a paragraph will be given, and then the MCQ questions based on it will be asked.
The above Case studies for Class 10 Maths will help you to boost your scores as Case Study questions have been coming in your examinations. These CBSE Class 10 Mathematics Case Studies have been developed by experienced teachers of cbseexpert.com for the benefit of Class 10 students.
- Class 10th Science Case Study Questions
- Assertion and Reason Questions of Class 10th Science
- Assertion and Reason Questions of Class 10th Social Science
Class 10 Maths Syllabus 2024
Chapter-1 real numbers.
Starting with an introduction to real numbers, properties of real numbers, Euclid’s division lemma, fundamentals of arithmetic, Euclid’s division algorithm, revisiting irrational numbers, revisiting rational numbers and their decimal expansions followed by a bunch of problems for a thorough and better understanding.
Chapter-2 Polynomials
This chapter is quite important and marks securing topics in the syllabus. As this chapter is repeated almost every year, students find this a very easy and simple subject to understand. Topics like the geometrical meaning of the zeroes of a polynomial, the relationship between zeroes and coefficients of a polynomial, division algorithm for polynomials followed with exercises and solved examples for thorough understanding.
Chapter-3 Pair of Linear Equations in Two Variables
This chapter is very intriguing and the topics covered here are explained very clearly and perfectly using examples and exercises for each topic. Starting with the introduction, pair of linear equations in two variables, graphical method of solution of a pair of linear equations, algebraic methods of solving a pair of linear equations, substitution method, elimination method, cross-multiplication method, equations reducible to a pair of linear equations in two variables, etc are a few topics that are discussed in this chapter.
Chapter-4 Quadratic Equations
The Quadratic Equations chapter is a very important and high priority subject in terms of examination, and securing as well as the problems are very simple and easy. Problems like finding the value of X from a given equation, comparing and solving two equations to find X, Y values, proving the given equation is quadratic or not by knowing the highest power, from the given statement deriving the required quadratic equation, etc are few topics covered in this chapter and also an ample set of problems are provided for better practice purposes.
Chapter-5 Arithmetic Progressions
This chapter is another interesting and simpler topic where the problems here are mostly based on a single formula and the rest are derivations of the original one. Beginning with a basic brief introduction, definitions of arithmetic progressions, nth term of an AP, the sum of first n terms of an AP are a few important and priority topics covered under this chapter. Apart from that, there are many problems and exercises followed with each topic for good understanding.
Chapter-6 Triangles
This chapter Triangle is an interesting and easy chapter and students often like this very much and a securing unit as well. Here beginning with the introduction to triangles followed by other topics like similar figures, the similarity of triangles, criteria for similarity of triangles, areas of similar triangles, Pythagoras theorem, along with a page summary for revision purposes are discussed in this chapter with examples and exercises for practice purposes.
Chapter-7 Coordinate Geometry
Here starting with a general introduction, distance formula, section formula, area of the triangle are a few topics covered in this chapter followed with examples and exercises for better and thorough practice purposes.
Chapter-8 Introduction to Trigonometry
As trigonometry is a very important and vast subject, this topic is divided into two parts where one chapter is Introduction to Trigonometry and another part is Applications of Trigonometry. This Introduction to Trigonometry chapter is started with a general introduction, trigonometric ratios, trigonometric ratios of some specific angles, trigonometric ratios of complementary angles, trigonometric identities, etc are a few important topics covered in this chapter.
Chapter-9 Applications of Trigonometry
This chapter is the continuation of the previous chapter, where the various modeled applications are discussed here with examples and exercises for better understanding. Topics like heights and distances are covered here and at the end, a summary is provided with all the important and frequently used formulas used in this chapter for solving the problems.
Chapter-10 Circle
Beginning with the introduction to circles, tangent to a circle, several tangents from a point on a circle are some of the important topics covered in this chapter. This chapter being practical, there are an ample number of problems and solved examples for better understanding and practice purposes.
Chapter-11 Constructions
This chapter has more practical problems than theory-based definitions. Beginning with a general introduction to constructions, tools used, etc, the topics like division of a line segment, construction of tangents to a circle, and followed with few solved examples that help in solving the exercises provided after each topic.
Chapter-12 Areas related to Circles
This chapter problem is exclusively formula based wherein topics like perimeter and area of a circle- A Review, areas of sector and segment of a circle, areas of combinations of plane figures, and a page summary is provided just as a revision of the topics and formulas covered in the entire chapter and also there are many exercises and solved examples for practice purposes.
Chapter-13 Surface Areas and Volumes
Starting with the introduction, the surface area of a combination of solids, the volume of a combination of solids, conversion of solid from one shape to another, frustum of a cone, etc are to name a few topics explained in detail provided with a set of examples for a better comprehension of the concepts.
Chapter-14 Statistics
In this chapter starting with an introduction, topics like mean of grouped data, mode of grouped data, a median of grouped, graphical representation of cumulative frequency distribution are explained in detail with exercises for practice purposes. This chapter being a simple and easy subject, securing the marks is not difficult for students.
Chapter-15 Probability
Probability is another simple and important chapter in examination point of view and as seeking knowledge purposes as well. Beginning with an introduction to probability, an important topic called A theoretical approach is explained here. Since this chapter is one of the smallest in the syllabus and problems are also quite easy, students often like this chapter
Leave a Comment Cancel reply
Save my name, email, and website in this browser for the next time I comment.
Download India's best Exam Preparation App Now.
Key Features
- Revision Notes
- Important Questions
- Previous Years Questions
- Case-Based Questions
- Assertion and Reason Questions
No thanks, I’m not interested!
- NCERT Solutions
- NCERT Solutions for Class 10
- NCERT Solutions for Class 10 Maths
- Chapter 6: Triangles
NCERT Solutions for Class 10 Maths Chapter 6 Triangles
Ncert solutions for class 10 maths chapter 6 – download free pdf.
NCERT Solutions for Class 10 Maths Chapter 6 Triangles are provided here, which is considered to be one of the most important study materials for the students studying in CBSE Class 10. Chapter 6 of NCERT Solutions for Class 10 Maths is well structured in accordance with the CBSE Syllabus for 2023-24. It covers a vast topic, including a number of rules and theorems. Students often tend to get confused about which theorem to use while solving a variety of questions.
Download Exclusively Curated Chapter Notes for Class 10 Maths Chapter – 6 Triangles
Download most important questions for class 10 maths chapter – 6 triangles.
The solutions provided at BYJU’S are designed in such a way that every step is explained clearly and in detail. The Solutions for NCERT Class 10 Maths are prepared by the subject experts to help students prepare better for their board exams. These solutions will be helpful not only for exam preparations, but also in solving homework and assignments.
The CBSE Class 10 examination often asks questions, either directly or indirectly, from the NCERT textbooks. Thus, the NCERT Solutions for Chapter 6 Triangles of Class 10 Maths is one of the best resources to prepare, and equip oneself to solve any type of questions in the exam, from the chapter. It is highly recommended that the students practise these NCERT Solutions on a regular basis to excel in the Class 10 board examinations.
- Chapter 1 Real Numbers
- Chapter 2 Polynomials
- Chapter 3 Pair of Linear Equations in Two Variables
- Chapter 4 Quadratic Equations
- Chapter 5 Arithmetic Progressions
- Chapter 6 Triangles
- Chapter 7 Coordinate Geometry
- Chapter 8 Introduction to Trigonometry
- Chapter 9 Some Applications of Trigonometry
- Chapter 10 Circles
- Chapter 11 Constructions
- Chapter 12 Areas Related to Circles
- Chapter 13 Surface Areas and Volumes
- Chapter 14 Statistics
- Chapter 15 Probability
carouselExampleControls112

Previous Next
Access Answers of NCERT Class 10 Maths Chapter 6 – Triangles
Exercise 6.1 page: 122.
1. Fill in the blanks using correct word given in the brackets:-
(i) All circles are __________. (congruent, similar)
Answer: Similar
(ii) All squares are __________. (similar, congruent)
(iii) All __________ triangles are similar. (isosceles, equilateral) Answer: Equilateral
(iv) Two polygons of the same number of sides are similar, if (a) their corresponding angles are __________ and (b) their corresponding sides are __________. (equal, proportional)
Answer: (a) Equal
(b) Proportional
2. Give two different examples of pair of (i) Similar figures (ii) Non-similar figures

3. State whether the following quadrilaterals are similar or not:

From the given two figures, we can see their corresponding angles are different or unequal. Therefore, they are not similar.
Exercise 6.2 Page: 128
1. In figure. (i) and (ii), DE || BC. Find EC in (i) and AD in (ii).

(i) Given, in △ ABC, DE∥BC
⇒1.5/3 = 1/EC
⇒EC = 3/1.5
EC = 3×10/15 = 2 cm
Hence, EC = 2 cm.
(ii) Given, in △ ABC, DE∥BC
⇒ AD/7.2 = 1.8 / 5.4
⇒ AD = 1.8 ×7.2/5.4 = (18/10)×(72/10)×(10/54) = 24/10
Hence, AD = 2.4 cm.
2. E and F are points on the sides PQ and PR, respectively of a ΔPQR. For each of the following cases, state whether EF || QR. (i) PE = 3.9 cm, EQ = 3 cm, PF = 3.6 cm and FR = 2.4 cm
(ii) PE = 4 cm, QE = 4.5 cm, PF = 8 cm and RF = 9 cm (iii) PQ = 1.28 cm, PR = 2.56 cm, PE = 0.18 cm and PF = 0.63 cm
Given, in ΔPQR, E and F are two points on side PQ and PR, respectively. See the figure below;

(i) Given, PE = 3.9 cm, EQ = 3 cm, PF = 3.6 cm and FR = 2,4 cm
Therefore, by using Basic proportionality theorem, we get,
PE/EQ = 3.9/3 = 39/30 = 13/10 = 1.3
And PF/FR = 3.6/2.4 = 36/24 = 3/2 = 1.5
So, we get, PE/EQ ≠ PF/FR
Hence, EF is not parallel to QR.
(ii) Given, PE = 4 cm, QE = 4.5 cm, PF = 8cm and RF = 9cm
PE/QE = 4/4.5 = 40/45 = 8/9
And, PF/RF = 8/9
So, we get here,
PE/QE = PF/RF
Hence, EF is parallel to QR.
(iii) Given, PQ = 1.28 cm, PR = 2.56 cm, PE = 0.18 cm and PF = 0.36 cm
From the figure,
EQ = PQ – PE = 1.28 – 0.18 = 1.10 cm
And, FR = PR – PF = 2.56 – 0.36 = 2.20 cm
So, PE/EQ = 0.18/1.10 = 18/110 = 9/55 …………. (i)
And, PE/FR = 0.36/2.20 = 36/220 = 9/55 ………… (ii)
PE/EQ = PF/FR
3. In the figure, if LM || CB and LN || CD, prove that AM/AB = AN/AD

In the given figure, we can see, LM || CB,
By using basic proportionality theorem, we get,
AM/AB = AL/AC ……………………..(i)
Similarly, given, LN || CD and using basic proportionality theorem,
∴AN/AD = AL/AC ……………………………(ii)
From equation (i) and (ii) , we get,
AM/AB = AN/AD
Hence, proved.
4. In the figure, DE||AC and DF||AE. Prove that BF/FE = BE/EC

In ΔABC, given as, DE || AC
Thus, by using Basic Proportionality Theorem, we get,
∴BD/DA = BE/EC ……………………………………………… (i)
In ΔBAE, given as, DF || AE
∴BD/DA = BF/FE ……………………………………………… (ii)
From equation (i) and (ii) , we get
BE/EC = BF/FE
5. In the figure, DE||OQ and DF||OR, show that EF||QR.

In ΔPQO, DE || OQ
So by using Basic Proportionality Theorem,
PD/DO = PE/EQ……………… ..(i)
Again given, in ΔPOR, DF || OR,
PD/DO = PF/FR………………… (ii)
Therefore, by converse of Basic Proportionality Theorem,
EF || QR, in ΔPQR.
6. In the figure, A, B and C are points on OP, OQ and OR respectively such that AB || PQ and AC || PR. Show that BC || QR.

Given here,
In ΔOPQ, AB || PQ
By using Basic Proportionality Theorem,
OA/AP = OB/BQ……………. (i)
Also given,
In ΔOPR, AC || PR
By using Basic Proportionality Theorem
∴ OA/AP = OC/CR……………(ii)
OB/BQ = OC/CR
In ΔOQR, BC || QR.
7. Using Basic proportionality theorem, prove that a line drawn through the mid-points of one side of a triangle parallel to another side bisects the third side. (Recall that you have proved it in Class IX).

Given, in ΔABC, D is the midpoint of AB such that AD=DB.
A line parallel to BC intersects AC at E as shown in above figure such that DE || BC.
We have to prove that E is the mid point of AC.
Since, D is the mid-point of AB.
⇒AD/DB = 1 …………………………. (i)
In ΔABC, DE || BC,
Therefore, AD/DB = AE/EC
From equation (i), we can write,
⇒ 1 = AE/EC
Hence, proved, E is the midpoint of AC.
8. Using Converse of basic proportionality theorem, prove that the line joining the mid-points of any two sides of a triangle is parallel to the third side. (Recall that you have done it in Class IX).
Given, in ΔABC, D and E are the mid points of AB and AC, respectively, such that,
AD=BD and AE=EC.

We have to prove that: DE || BC.
Since, D is the midpoint of AB
⇒AD/BD = 1……………………………….. (i)
Also given, E is the mid-point of AC.
⇒ AE/EC = 1
AD/BD = AE/EC
By converse of Basic Proportionality Theorem,
9. ABCD is a trapezium in which AB || DC and its diagonals intersect each other at the point O. Show that AO/BO = CO/DO.
Given, ABCD is a trapezium where AB || DC and diagonals AC and BD intersect each other at O.

We have to prove, AO/BO = CO/DO
From the point O, draw a line EO touching AD at E, in such a way that,
EO || DC || AB
In ΔADC, we have OE || DC
Therefore, by using Basic Proportionality Theorem
AE/ED = AO/CO ……………..(i)
Now, In ΔABD, OE || AB
DE/EA = DO/BO …………….(ii)
AO/CO = BO/DO
⇒AO/BO = CO/DO
10. The diagonals of a quadrilateral ABCD intersect each other at the point O such that AO/BO = CO/DO. Show that ABCD is a trapezium.
Given, Quadrilateral ABCD where AC and BD intersect each other at O such that,
AO/BO = CO/DO.

We have to prove here, ABCD is a trapezium
In ΔDAB, EO || AB
DE/EA = DO/OB ……………………(i)
Also, given,
AO/BO = CO/DO
⇒ AO/CO = BO/DO
⇒ CO/AO = DO/BO
⇒DO/OB = CO/AO …………………………..(ii)
DE/EA = CO/AO
Therefore, by using converse of Basic Proportionality Theorem,
EO || DC also EO || AB
⇒ AB || DC.
Hence, quadrilateral ABCD is a trapezium with AB || CD.
Exercise 6.3 Page: 138
1. State which pairs of triangles in the figure are similar. Write the similarity criterion used by you for answering the question and also write the pairs of similar triangles in the symbolic form:

(i) Given, in ΔABC and ΔPQR,
∠A = ∠P = 60°
∠B = ∠Q = 80°
∠C = ∠R = 40°
Therefore, by AAA similarity criterion,
∴ ΔABC ~ ΔPQR
(ii) Given, in ΔABC and ΔPQR,
AB/QR = 2/4 = 1/2,
BC/RP = 2.5/5 = 1/2,
CA/PA = 3/6 = 1/2
By SSS similarity criterion,
ΔABC ~ ΔQRP
(iii) Given, in ΔLMP and ΔDEF,
LM = 2.7, MP = 2, LP = 3, EF = 5, DE = 4, DF = 6
MP/DE = 2/4 = 1/2
PL/DF = 3/6 = 1/2
LM/EF = 2.7/5 = 27/50
Here , MP/DE = PL/DF ≠ LM/EF
Therefore, ΔLMP and ΔDEF are not similar.
(iv) In ΔMNL and ΔQPR, it is given,
MN/QP = ML/QR = 1/2
∠M = ∠Q = 70°
Therefore, by SAS similarity criterion
∴ ΔMNL ~ ΔQPR
(v) In ΔABC and ΔDEF, given that,
AB = 2.5, BC = 3, ∠A = 80°, EF = 6, DF = 5, ∠F = 80°
Here , AB/DF = 2.5/5 = 1/2
And, BC/EF = 3/6 = 1/2
Hence, ΔABC and ΔDEF are not similar.
(vi) In ΔDEF, by sum of angles of triangles, we know that,
∠D + ∠E + ∠F = 180°
⇒ 70° + 80° + ∠F = 180°
⇒ ∠F = 180° – 70° – 80°
Similarly, In ΔPQR,
∠P + ∠Q + ∠R = 180 (Sum of angles of Δ)
⇒ ∠P + 80° + 30° = 180°
⇒ ∠P = 180° – 80° -30°
Now, comparing both the triangles, ΔDEF and ΔPQR, we have
∠D = ∠P = 70°
∠F = ∠Q = 80°
∠F = ∠R = 30°
Hence, ΔDEF ~ ΔPQR
2. In figure 6.35, ΔODC ~ ΔOBA, ∠ BOC = 125° and ∠ CDO = 70°. Find ∠ DOC, ∠ DCO and ∠ OAB.

As we can see from the figure, DOB is a straight line.
Therefore, ∠DOC + ∠ COB = 180°
⇒ ∠DOC = 180° – 125° (Given, ∠ BOC = 125°)
In ΔDOC, sum of the measures of the angles of a triangle is 180º
Therefore, ∠DCO + ∠ CDO + ∠ DOC = 180°
⇒ ∠DCO + 70º + 55º = 180°(Given, ∠ CDO = 70°)
⇒ ∠DCO = 55°
It is given that, ΔODC ~ ΔOBA,
Therefore, ΔODC ~ ΔOBA.
Hence, corresponding angles are equal in similar triangles
∠OAB = ∠OCD
⇒ ∠ OAB = 55°
⇒ ∠OAB = 55°
3. Diagonals AC and BD of a trapezium ABCD with AB || DC intersect each other at the point O. Using a similarity criterion for two triangles, show that AO/OC = OB/OD

In ΔDOC and ΔBOA,
AB || CD, thus alternate interior angles will be equal,
∴∠CDO = ∠ABO
∠DCO = ∠BAO
Also, for the two triangles ΔDOC and ΔBOA, vertically opposite angles will be equal;
∴∠DOC = ∠BOA
Hence, by AAA similarity criterion,
ΔDOC ~ ΔBOA
Thus, the corresponding sides are proportional.
DO/BO = OC/OA
⇒OA/OC = OB/OD
4. In the fig.6.36, QR/QS = QT/PR and ∠1 = ∠2. Show that ΔPQS ~ ΔTQR.

∠PQR = ∠PRQ
∴ PQ = PR ……………………… (i)
QR/QS = QT/PRUsing equation (i) , we get
QR/QS = QT/QP ……………….(ii)
In ΔPQS and ΔTQR, by equation (ii),
QR/QS = QT/QP
5. S and T are point on sides PR and QR of ΔPQR such that ∠P = ∠RTS. Show that ΔRPQ ~ ΔRTS.
Given, S and T are point on sides PR and QR of ΔPQR
And ∠P = ∠RTS.

In ΔRPQ and ΔRTS,
∠RTS = ∠QPS (Given)
∠R = ∠R (Common angle)
∴ ΔRPQ ~ ΔRTS (AA similarity criterion)
6. In the figure, if ΔABE ≅ ΔACD, show that ΔADE ~ ΔABC.

Given, ΔABE ≅ ΔACD.
∴ AB = AC [By CPCT] ………………………………. (i)
And, AD = AE [By CPCT] …………………………… (ii)
In ΔADE and ΔABC, dividing eq.(ii) by eq(i),
AD/AB = AE/AC
7. In the figure, altitudes AD and CE of ΔABC intersect each other at the point P. Show that:

(i) ΔAEP ~ ΔCDP (ii) ΔABD ~ ΔCBE (iii) ΔAEP ~ ΔADB (iv) ΔPDC ~ ΔBEC
Given, altitudes AD and CE of ΔABC intersect each other at the point P.
(i) In ΔAEP and ΔCDP,
∠AEP = ∠CDP (90° each)
∠APE = ∠CPD (Vertically opposite angles)
Hence, by AA similarity criterion,
ΔAEP ~ ΔCDP
(ii) In ΔABD and ΔCBE,
∠ADB = ∠CEB ( 90° each)
∠ABD = ∠CBE (Common Angles)
ΔABD ~ ΔCBE
(iii) In ΔAEP and ΔADB,
∠AEP = ∠ADB (90° each)
∠PAE = ∠DAB (Common Angles)
ΔAEP ~ ΔADB
(iv) In ΔPDC and ΔBEC,
∠PDC = ∠BEC (90° each)
∠PCD = ∠BCE (Common angles)
ΔPDC ~ ΔBEC
8. E is a point on the side AD produced of a parallelogram ABCD and BE intersects CD at F. Show that ΔABE ~ ΔCFB.
Given, E is a point on the side AD produced of a parallelogram ABCD and BE intersects CD at F. Consider the figure below,

In ΔABE and ΔCFB,
∠A = ∠C (Opposite angles of a parallelogram)
∠AEB = ∠CBF (Alternate interior angles as AE || BC)
∴ ΔABE ~ ΔCFB (AA similarity criterion)
9. In the figure, ABC and AMP are two right triangles, right angled at B and M, respectively, prove that:

(i) ΔABC ~ ΔAMP
(ii) CA/PA = BC/MP
Given, ABC and AMP are two right triangles, right angled at B and M, respectively.
(i) In ΔABC and ΔAMP, we have,
∠CAB = ∠MAP (common angles)
∠ABC = ∠AMP = 90° (each 90°)
∴ ΔABC ~ ΔAMP (AA similarity criterion)
(ii) As, ΔABC ~ ΔAMP (AA similarity criterion)
If two triangles are similar then the corresponding sides are always equal,
Hence, CA/PA = BC/MP
10. CD and GH are respectively the bisectors of ∠ACB and ∠EGF such that D and H lie on sides AB and FE of ΔABC and ΔEFG respectively. If ΔABC ~ ΔFEG, Show that:
(i) CD/GH = AC/FG (ii) ΔDCB ~ ΔHGE (iii) ΔDCA ~ ΔHGF
Given, CD and GH are respectively the bisectors of ∠ACB and ∠EGF such that D and H lie on sides AB and FE of ΔABC and ΔEFG, respectively.

(i) From the given condition,
ΔABC ~ ΔFEG.
∴ ∠A = ∠F, ∠B = ∠E, and ∠ACB = ∠FGE
Since, ∠ACB = ∠FGE
∴ ∠ACD = ∠FGH (Angle bisector)
And, ∠DCB = ∠HGE (Angle bisector)
In ΔACD and ΔFGH,
∠ACD = ∠FGH
∴ ΔACD ~ ΔFGH (AA similarity criterion)
⇒CD/GH = AC/FG
(ii) In ΔDCB and ΔHGE,
∠DCB = ∠HGE (Already proved)
∠B = ∠E (Already proved)
∴ ΔDCB ~ ΔHGE (AA similarity criterion)
(iii) In ΔDCA and ΔHGF,
∠ACD = ∠FGH (Already proved)
∠A = ∠F (Already proved)
∴ ΔDCA ~ ΔHGF (AA similarity criterion)
11. In the following figure, E is a point on side CB produced of an isosceles triangle ABC with AB = AC. If AD ⊥ BC and EF ⊥ AC, prove that ΔABD ~ ΔECF.

Given, ABC is an isosceles triangle.
⇒ ∠ABD = ∠ECF
In ΔABD and ΔECF,
∠ADB = ∠EFC (Each 90°)
∠BAD = ∠CEF (Already proved)
∴ ΔABD ~ ΔECF (using AA similarity criterion)
12. Sides AB and BC and median AD of a triangle ABC are respectively proportional to sides PQ and QR and median PM of ΔPQR (see Fig 6.41). Show that ΔABC ~ ΔPQR.

Given, ΔABC and ΔPQR, AB, BC and median AD of ΔABC are proportional to sides PQ, QR and median PM of ΔPQR
i.e. AB/PQ = BC/QR = AD/PM
We have to prove: ΔABC ~ ΔPQR
As we know here,
AB/PQ = BC/QR = AD/PM

⇒AB/PQ = BC/QR = AD/PM (D is the midpoint of BC. M is the midpoint of QR)
⇒ ∠ABC = ∠PQR
In ΔABC and ΔPQR
AB/PQ = BC/QR …………………………. (i)
∠ABC = ∠PQR …………………………… (ii)
13. D is a point on the side BC of a triangle ABC such that ∠ADC = ∠BAC. Show that CA 2 = CB.CD
Given, D is a point on the side BC of a triangle ABC such that ∠ADC = ∠BAC.

In ΔADC and ΔBAC,
∠ADC = ∠BAC (Already given)
∠ACD = ∠BCA (Common angles)
∴ ΔADC ~ ΔBAC (AA similarity criterion)
We know that corresponding sides of similar triangles are in proportion.
∴ CA/CB = CD/CA
⇒ CA 2 = CB.CD.
14. Sides AB and AC and median AD of a triangle ABC are respectively proportional to sides PQ and PR and median PM of another triangle PQR. Show that ΔABC ~ ΔPQR.
Given: Two triangles ΔABC and ΔPQR in which AD and PM are medians such that;
AB/PQ = AC/PR = AD/PM
We have to prove, ΔABC ~ ΔPQR
Let us construct first: Produce AD to E so that AD = DE. Join CE, Similarly produce PM to N such that PM = MN, also Join RN.

In ΔABD and ΔCDE, we have
⇒ AB = CE [By CPCT] ………………………….. (i)
Also, in ΔPQM and ΔMNR,
⇒ PQ = RN [CPCT] ……………………………… (ii)
Now, AB/PQ = AC/PR = AD/PM
From equation (i) and (ii) ,
⇒CE/RN = AC/PR = AD/PM
⇒ CE/RN = AC/PR = 2AD/2PM
Therefore, ∠2 = ∠4
Similarly, ∠1 = ∠3
∴ ∠1 + ∠2 = ∠3 + ∠4
⇒ ∠A = ∠P ……………………………………………. (iii)
Now, in ΔABC and ΔPQR, we have
AB/PQ = AC/PR (Already given)
From equation (iii),
15. A vertical pole of a length 6 m casts a shadow 4m long on the ground and at the same time a tower casts a shadow 28 m long. Find the height of the tower.
Given, Length of the vertical pole = 6m
Shadow of the pole = 4 m
Let Height of tower = h m
Length of shadow of the tower = 28 m

In ΔABC and ΔDEF,
∠C = ∠E (angular elevation of sum)
∠B = ∠F = 90°
∴ ΔABC ~ ΔDEF (AA similarity criterion)
∴ AB/DF = BC/EF (If two triangles are similar corresponding sides are proportional)
∴ 6/h = 4/28
⇒h = (6×28)/4
⇒ h = 6 × 7
⇒ h = 42 m
Hence, the height of the tower is 42 m.
16. If AD and PM are medians of triangles ABC and PQR, respectively where ΔABC ~ ΔPQR prove that AB/PQ = AD/PM.
Given, ΔABC ~ ΔPQR

We know that the corresponding sides of similar triangles are in proportion.
∴AB/PQ = AC/PR = BC/QR ……………………………(i )
Also, ∠A = ∠P, ∠B = ∠Q, ∠C = ∠R ………….….. (ii)
Since AD and PM are medians, they will divide their opposite sides.
∴ BD = BC/2 and QM = QR/2 ……………..…………. (iii)
From equations (i) and (iii) , we get
AB/PQ = BD/QM ……………………….(iv)
In ΔABD and ΔPQM,
From equation (ii), we have
From equation (iv), we have,
AB/PQ = BD/QM
∴ ΔABD ~ ΔPQM (SAS similarity criterion)
⇒AB/PQ = BD/QM = AD/PM
Exercise 6.4 Page: 143
1. Let ΔABC ~ ΔDEF and their areas be, respectively, 64 cm 2 and 121 cm 2 . If EF = 15.4 cm, find BC.
Solution: Given, ΔABC ~ ΔDEF,
Area of ΔABC = 64 cm 2
Area of ΔDEF = 121 cm 2
EF = 15.4 cm

As we know, if two triangles are similar, ratio of their areas are equal to the square of the ratio of their corresponding sides,
= AC 2 /DF 2 = BC 2 /EF 2
∴ 64/121 = BC 2 /EF 2
⇒ (8/11) 2 = (BC/15.4) 2
⇒ 8/11 = BC/15.4
⇒ BC = 8×15.4/11
⇒ BC = 8 × 1.4
⇒ BC = 11.2 cm
2. Diagonals of a trapezium ABCD with AB || DC intersect each other at the point O. If AB = 2CD, find the ratio of the areas of triangles AOB and COD.
Given, ABCD is a trapezium with AB || DC. Diagonals AC and BD intersect each other at point O.

In ΔAOB and ΔCOD, we have
∠1 = ∠2 (Alternate angles)
∠3 = ∠4 (Alternate angles)
∠5 = ∠6 (Vertically opposite angle)
As we know, If two triangles are similar then the ratio of their areas are equal to the square of the ratio of their corresponding sides. Therefore,
Area of (ΔAOB)/Area of (ΔCOD) = AB 2 /CD 2
∴ Area of (ΔAOB)/Area of (ΔCOD)
= 4CD 2 /CD 2 = 4/1
Hence, the required ratio of the area of ΔAOB and ΔCOD = 4:1
3. In the figure, ABC and DBC are two triangles on the same base BC. If AD intersects BC at O, show that area (ΔABC)/area (ΔDBC) = AO/DO.

Given, ABC and DBC are two triangles on the same base BC. AD intersects BC at O.
We have to prove: Area (ΔABC)/Area (ΔDBC) = AO/DO
Let us draw two perpendiculars AP and DM on line BC.

We know that area of a triangle = 1/2 × Base × Height

In ΔAPO and ΔDMO,
∠APO = ∠DMO (Each 90°)
∠AOP = ∠DOM (Vertically opposite angles)
∴ ΔAPO ~ ΔDMO (AA similarity criterion)
∴ AP/DM = AO/DO
⇒ Area (ΔABC)/Area (ΔDBC) = AO/DO.
4. If the areas of two similar triangles are equal, prove that they are congruent.
Say ΔABC and ΔPQR are two similar triangles and equal in area

Now let us prove ΔABC ≅ ΔPQR.
Since, ΔABC ~ ΔPQR
∴ Area of (ΔABC)/Area of (ΔPQR) = BC 2 /QR 2
⇒ BC 2 /QR 2 =1 [Since, Area(ΔABC) = (ΔPQR)
⇒ BC 2 /QR 2
Similarly, we can prove that
AB = PQ and AC = PR
5. D, E and F are respectively the mid-points of sides AB, BC and CA of ΔABC. Find the ratio of the area of ΔDEF and ΔABC.

6. Prove that the ratio of the areas of two similar triangles is equal to the square of the ratio of their corresponding medians.
Given: AM and DN are the medians of triangles ABC and DEF respectively and ΔABC ~ ΔDEF.

We have to prove: Area(ΔABC)/Area(ΔDEF) = AM 2 /DN 2
Since, ΔABC ~ ΔDEF (Given)
∴ Area(ΔABC)/Area(ΔDEF) = (AB 2 /DE 2 ) …………………………… (i)
and, AB/DE = BC/EF = CA/FD ……………………………………… (ii)

In ΔABM and ΔDEN,
Since ΔABC ~ ΔDEF
⇒ AB/DE = AM/DN ………………………………………………….. (iii)
∴ ΔABM ~ ΔDEN
As the areas of two similar triangles are proportional to the squares of the corresponding sides.
∴ area(ΔABC)/area(ΔDEF) = AB 2 /DE 2 = AM 2 /DN 2
7. Prove that the area of an equilateral triangle described on one side of a square is equal to half the area of the equilateral triangle described on one of its diagonals.

Area(ΔBQC) = ½ Area(ΔAPC)
Since, ΔAPC and ΔBQC are both equilateral triangles, as per given,
∴ area(ΔAPC)/area(ΔBQC) = (AC 2 /BC 2 ) = AC 2 /BC 2
Since, Diagonal = √2 side = √2 BC = AC

⇒ area(ΔAPC) = 2 × area(ΔBQC)
⇒ area(ΔBQC) = 1/2area(ΔAPC)
Tick the correct answer and justify:
8. ABC and BDE are two equilateral triangles such that D is the mid-point of BC. Ratio of the area of triangles ABC and BDE is (A) 2 : 1 (B) 1 : 2 (C) 4 : 1 (D) 1 : 4
Given , ΔABC and ΔBDE are two equilateral triangle. D is the midpoint of BC.

∴ BD = DC = 1/2BC
Let each side of triangle is 2 a .
As, ΔABC ~ ΔBDE
∴ Area(ΔABC)/Area(ΔBDE) = AB 2 /BD 2 = (2 a ) 2 /( a ) 2 = 4 a 2 / a 2 = 4/1 = 4:1
Hence, the correct answer is (C).
9. Sides of two similar triangles are in the ratio 4 : 9. Areas of these triangles are in the ratio (A) 2 : 3 (B) 4 : 9 (C) 81 : 16 (D) 16 : 81
Given, Sides of two similar triangles are in the ratio 4 : 9.

Let ABC and DEF are two similar triangles, such that,
ΔABC ~ ΔDEF
And AB/DE = AC/DF = BC/EF = 4/9
As, the ratio of the areas of these triangles will be equal to the square of the ratio of the corresponding sides,
∴ Area(ΔABC)/Area(ΔDEF) = AB 2 /DE 2
∴ Area(ΔABC)/Area(ΔDEF) = (4/9) 2 = 16/81 = 16:81
Hence, the correct answer is (D).
Exercise 6.5 Page: 150
1. Sides of triangles are given below. Determine which of them are right triangles. In case of a right triangle, write the length of its hypotenuse.
(i) 7 cm, 24 cm, 25 cm (ii) 3 cm, 8 cm, 6 cm (iii) 50 cm, 80 cm, 100 cm (iv) 13 cm, 12 cm, 5 cm
(i) Given, sides of the triangle are 7 cm, 24 cm, and 25 cm.
Squaring the lengths of the sides of the, we will get 49, 576, and 625.
49 + 576 = 625
(7) 2 + (24) 2 = (25) 2
Therefore, the above equation satisfies, Pythagoras theorem. Hence, it is right angled triangle.
Length of Hypotenuse = 25 cm
(ii) Given, sides of the triangle are 3 cm, 8 cm, and 6 cm.
Squaring the lengths of these sides, we will get 9, 64, and 36.
Clearly, 9 + 36 ≠ 64
Or, 3 2 + 6 2 ≠ 8 2
Therefore, the sum of the squares of the lengths of two sides is not equal to the square of the length of the hypotenuse.
Hence, the given triangle does not satisfies Pythagoras theorem.
(iii) Given, sides of triangle’s are 50 cm, 80 cm, and 100 cm.
Squaring the lengths of these sides, we will get 2500, 6400, and 10000.
However, 2500 + 6400 ≠ 10000
Or, 50 2 + 80 2 ≠ 100 2
As you can see, the sum of the squares of the lengths of two sides is not equal to the square of the length of the third side.
Therefore, the given triangle does not satisfies Pythagoras theorem.
Hence, it is not a right triangle.
(iv) Given, sides are 13 cm, 12 cm, and 5 cm.
Squaring the lengths of these sides, we will get 169, 144, and 25.
Thus, 144 +25 = 169
Or, 12 2 + 5 2 = 13 2
The sides of the given triangle are satisfying Pythagoras theorem.
Therefore, it is a right triangle.
Hence, length of the hypotenuse of this triangle is 13 cm.
2. PQR is a triangle right angled at P and M is a point on QR such that PM ⊥ QR. Show that PM 2 = QM × MR.
Given, ΔPQR is right angled at P is a point on QR such that PM ⊥QR

We have to prove, PM 2 = QM × MR
In ΔPQM, by Pythagoras theorem
PQ 2 = PM 2 + QM 2
Or, PM 2 = PQ 2 – QM 2 …………………………….. (i)
In ΔPMR, by Pythagoras theorem
PR 2 = PM 2 + MR 2
Or, PM 2 = PR 2 – MR 2 ……………………………………….. (ii)
Adding equation, (i) and (ii) , we get,
2PM 2 = (PQ 2 + PM 2 ) – (QM 2 + MR 2 )
= (QM + MR) 2 – QM 2 – MR 2
∴ PM 2 = QM × MR
3. In Figure, ABD is a triangle right angled at A and AC ⊥ BD. Show that (i) AB 2 = BC × BD (ii) AC 2 = BC × DC (iii) AD 2 = BD × CD

(i) In ΔADB and ΔCAB,
∠DAB = ∠ACB (Each 90°)
∠ABD = ∠CBA (Common angles)
⇒ AB/CB = BD/AB
⇒ AB 2 = CB × BD
(ii) Let ∠CAB = x
∠CBA = 180° – 90° – x
∠CBA = 90° – x
Similarly, in ΔCAD
∠CAD = 90° – ∠CBA
= 90° – x
∠CDA = 180° – 90° – (90° – x)
In ΔCBA and ΔCAD, we have
∠CBA = ∠CAD
∠CAB = ∠CDA
∠ACB = ∠DCA (Each 90°)
⇒ AC/DC = BC/AC
⇒ AC 2 = DC × BC
(iii) In ΔDCA and ΔDAB,
∠DCA = ∠DAB (Each 90°)
∠CDA = ∠ADB (common angles)
⇒ DC/DA = DA/DA
⇒ AD 2 = BD × CD
4. ABC is an isosceles triangle right angled at C. Prove that AB 2 = 2AC 2 .
Given, ΔABC is an isosceles triangle right angled at C.

In ΔACB, ∠C = 90°
AC = BC (By isosceles triangle property)
AB 2 = 2AC 2
5. ABC is an isosceles triangle with AC = BC. If AB 2 = 2AC 2 , prove that ABC is a right triangle.
Given, ΔABC is an isosceles triangle having AC = BC and AB 2 = 2AC 2

AB 2 = AC 2 + AC 2
Hence, by Pythagoras theorem ΔABC is right angle triangle.
6. ABC is an equilateral triangle of side 2a. Find each of its altitudes .
Given, ABC is an equilateral triangle of side 2a.

Draw, AD ⊥ BC
In ΔADB and ΔADC,
Therefore, ΔADB ≅ ΔADC by RHS congruence.
In right angled ΔADB,
AB 2 = AD 2 + BD 2
(2 a ) 2 = AD 2 + a 2
⇒ AD 2 = 4 a 2 – a 2
⇒ AD 2 = 3 a 2
⇒ AD = √3a
7. Prove that the sum of the squares of the sides of rhombus is equal to the sum of the squares of its diagonals.
Given, ABCD is a rhombus whose diagonals AC and BD intersect at O.

We have to prove, as per the question,
AB 2 + BC 2 + CD 2 + AD 2 = AC 2 + BD 2
Since, the diagonals of a rhombus bisect each other at right angles.
Therefore, AO = CO and BO = DO
AD 2 = AO 2 + DO 2 …………………….. (ii)
DC 2 = DO 2 + CO 2 …………………….. (iii)
BC 2 = CO 2 + BO 2 …………………….. (iv)
Adding equations (i) + (ii) + (iii) + (iv) , we get,
AB 2 + AD 2 + DC 2 + BC 2 = 2(AO 2 + BO 2 + DO 2 + CO 2 )
= (2AO) 2 + (2BO) 2 = AC 2 + BD 2
AB 2 + AD 2 + DC 2 + BC 2 = AC 2 + BD 2
8. In Fig. 6.54, O is a point in the interior of a triangle.

ABC, OD ⊥ BC, OE ⊥ AC and OF ⊥ AB. Show that: (i) OA 2 + OB 2 + OC 2 – OD 2 – OE 2 – OF 2 = AF 2 + BD 2 + CE 2 , (ii) AF 2 + BD 2 + CE 2 = AE 2 + CD 2 + BF 2 .
Given, in ΔABC, O is a point in the interior of a triangle.
And OD ⊥ BC, OE ⊥ AC and OF ⊥ AB.
Join OA, OB and OC

(i) By Pythagoras theorem in ΔAOF, we have
OA 2 = OF 2 + AF 2
Similarly, in ΔBOD
OB 2 = OD 2 + BD 2
Similarly, in ΔCOE
OC 2 = OE 2 + EC 2
Adding these equations,
OA 2 + OB 2 + OC 2 = OF 2 + AF 2 + OD 2 + BD 2 + OE 2 + EC 2
OA 2 + OB 2 + OC 2 – OD 2 – OE 2 – OF 2 = AF 2 + BD 2 + CE 2 .
(ii) AF 2 + BD 2 + EC 2 = (OA 2 – OE 2 ) + (OC 2 – OD 2 ) + (OB 2 – OF 2 )
∴ AF 2 + BD 2 + CE 2 = AE 2 + CD 2 + BF 2 .
9. A ladder 10 m long reaches a window 8 m above the ground. Find the distance of the foot of the ladder from base of the wall.
Given, a ladder 10 m long reaches a window 8 m above the ground.

Let BA be the wall and AC be the ladder,
Therefore, by Pythagoras theorem,
AC 2 = AB 2 + BC 2
10 2 = 8 2 + BC 2
BC 2 = 100 – 64
Therefore, the distance of the foot of the ladder from the base of the wall is 6 m.
10. A guy wire attached to a vertical pole of height 18 m is 24 m long and has a stake attached to the other end. How far from the base of the pole should the stake be driven so that the wire will be taut?
Given, a guy wire attached to a vertical pole of height 18 m is 24 m long and has a stake attached to the other end.

Let AB be the pole and AC be the wire.
By Pythagoras theorem,
24 2 = 18 2 + BC 2
BC 2 = 576 – 324
BC 2 = 252
BC = 6√7m
Therefore, the distance from the base is 6√7m.

Speed of first aeroplane = 1000 km/hr

Speed of second aeroplane = 1200 km/hr

In right angle ΔAOB, by Pythagoras Theorem,
AB 2 = AO 2 + OB 2
⇒ AB 2 = (1500) 2 + (1800) 2
⇒ AB = √(2250000 + 3240000)
⇒ AB = 300√61 km
Hence, the distance between two aeroplanes will be 300√61 km.
12. Two poles of heights 6 m and 11 m stand on a plane ground. If the distance between the feet of the poles is 12 m, find the distance between their tops.
Given, Two poles of heights 6 m and 11 m stand on a plane ground.
And distance between the feet of the poles is 12 m.

Let AB and CD be the poles of height 6m and 11m.
Therefore, CP = 11 – 6 = 5m
From the figure, it can be observed that AP = 12m
By Pythagoras theorem for ΔAPC, we get,
AP 2 = PC 2 + AC 2
(12m) 2 + (5m) 2 = (AC) 2
AC 2 = (144+25) m 2 = 169 m 2
Therefore, the distance between their tops is 13 m.
13. D and E are points on the sides CA and CB respectively of a triangle ABC right angled at C. Prove that AE 2 + BD 2 = AB 2 + DE 2 .
Given, D and E are points on the sides CA and CB respectively of a triangle ABC right angled at C.

By Pythagoras theorem in ΔACE, we get
AC 2 + CE 2 = AE 2 …………………………………………. (i)
In ΔBCD, by Pythagoras theorem, we get
BC 2 + CD 2 = BD 2 ……………………………….. (ii)
From equations (i) and (ii) , we get,
AC 2 + CE 2 + BC 2 + CD 2 = AE 2 + BD 2 ………….. (iii)
In ΔCDE, by Pythagoras theorem, we get
DE 2 = CD 2 + CE 2
In ΔABC, by Pythagoras theorem, we get
AB 2 = AC 2 + CB 2
Putting the above two values in equation (iii) , we get
DE 2 + AB 2 = AE 2 + BD 2 .
14. The perpendicular from A on side BC of a Δ ABC intersects BC at D such that DB = 3CD (see Figure). Prove that 2AB 2 = 2AC 2 + BC 2 .

Given, the perpendicular from A on side BC of a Δ ABC intersects BC at D such that;
AD ⊥BC and BD = 3CD
In right angle triangle, ADB and ADC, by Pythagoras theorem,
AB 2 = AD 2 + BD 2 ………………………. (i)
AC 2 = AD 2 + DC 2 …………………………….. (ii)
Subtracting equation (ii) from equation (i) , we get
AB 2 – AC 2 = BD 2 – DC 2
Therefore, AB 2 – AC 2 = BC 2 /2
⇒ 2(AB 2 – AC 2 ) = BC 2
⇒ 2AB 2 – 2AC 2 = BC 2
∴ 2AB 2 = 2AC 2 + BC 2 .
15. In an equilateral triangle ABC, D is a point on side BC such that BD = 1/3BC. Prove that 9AD 2 = 7AB 2 .
Given, ABC is an equilateral triangle.
And D is a point on side BC such that BD = 1/3BC

Let the side of the equilateral triangle be a , and AE be the altitude of ΔABC.
∴ BE = EC = BC/2 = a/2
And, AE = a√3/2
Given, BD = 1/3BC
DE = BE – BD = a/2 – a/3 = a/6
In ΔADE, by Pythagoras theorem,
AD 2 = AE 2 + DE 2

⇒ 9 AD 2 = 7 AB 2
16. In an equilateral triangle, prove that three times the square of one side is equal to four times the square of one of its altitudes.
Given, an equilateral triangle say ABC,

Let the sides of the equilateral triangle be of length a, and AE be the altitude of ΔABC.
∴ BE = EC = BC/2 = a/2
In ΔABE, by Pythagoras Theorem, we get
AB 2 = AE 2 + BE 2

4AE 2 = 3a 2
⇒ 4 × (Square of altitude) = 3 × (Square of one side)
17. Tick the correct answer and justify: In ΔABC, AB = 6√3 cm, AC = 12 cm and BC = 6 cm. The angle B is: (A) 120°
(B) 60° (C) 90°
Given, in ΔABC, AB = 6√3 cm, AC = 12 cm and BC = 6 cm.

We can observe that,
AB 2 = 108
AC 2 = 144
And, BC 2 = 36
AB 2 + BC 2 = AC 2
The given triangle, ΔABC, is satisfying Pythagoras theorem.
Therefore, the triangle is a right triangle, right-angled at B.
Exercise 6.6 Page: 152
1. In Figure, PS is the bisector of ∠ QPR of ∆ PQR. Prove that QS/PQ = SR/PR

Let us draw a line segment RT parallel to SP which intersects extended line segment QP at point T.
Given, PS is the angle bisector of ∠QPR. Therefore,
∠QPS = ∠SPR………………………………..(i)

As per the constructed figure,
∠SPR=∠PRT(Since, PS||TR)……………(ii)
∠QPS = ∠QRT(Since, PS||TR) …………..(iii)
From the above equations, we get,
In △QTR, by basic proportionality theorem,
QS/SR = QP/PT
Since, PT=TR
QS/SR = PQ/PR

- Let us join Point D and B.

BD ⊥AC, DM ⊥ BC and DN ⊥ AB
Now from the figure we have,
DN || CB, DM || AB and ∠B = 90 °
Therefore, DMBN is a rectangle.
So, DN = MB and DM = NB
The given condition which we have to prove, is when D is the foot of the perpendicular drawn from B to AC.
∴ ∠CDB = 90° ⇒ ∠2 + ∠3 = 90° ……………………. (i)
In ∆CDM, ∠1 + ∠2 + ∠DMC = 180°
⇒ ∠1 + ∠2 = 90° …………………………………….. (ii)
In ∆DMB, ∠3 + ∠DMB + ∠4 = 180°
⇒ ∠3 + ∠4 = 90° …………………………………….. (iii)
From equation (i) and (ii), we get
From equation (i) and (iii), we get
In ∆DCM and ∆BDM,
∠1 = ∠3 (Already Proved)
∠2 = ∠4 (Already Proved)
∴ ∆DCM ∼ ∆BDM (AA similarity criterion)
BM/DM = DM/MC
DN/DM = DM/MC (BM = DN)
⇒ DM 2 = DN × MC
(ii) In right triangle DBN,
∠5 + ∠7 = 90° ……………….. (iv)
In right triangle DAN,
∠6 + ∠8 = 90° ………………… (v)
D is the point in triangle, which is foot of the perpendicular drawn from B to AC.
∴ ∠ADB = 90° ⇒ ∠5 + ∠6 = 90° ………….. (vi)
From equation (iv) and (vi), we get,
From equation (v) and (vi), we get,
In ∆DNA and ∆BND,
∠6 = ∠7 (Already proved)
∠8 = ∠5 (Already proved)
∴ ∆DNA ∼ ∆BND (AA similarity criterion)
AN/DN = DN/NB
⇒ DN 2 = AN × NB
⇒ DN 2 = AN × DM (Since, NB = DM)
3. In Figure, ABC is a triangle in which ∠ABC > 90° and AD ⊥ CB produced. Prove that
AC 2 = AB 2 + BC 2 + 2 BC.BD.

By applying Pythagoras Theorem in ∆ADB, we get,
AB 2 = AD 2 + DB 2 ……………………… (i)
Again, by applying Pythagoras Theorem in ∆ACD, we get,
AC 2 = AD 2 + DC 2
AC 2 = AD 2 + (DB + BC) 2
AC 2 = AD 2 + DB 2 + BC 2 + 2DB × BC
AC 2 = AB 2 + BC 2 + 2DB × BC
4. In Figure, ABC is a triangle in which ∠ ABC < 90° and AD ⊥ BC. Prove that
AC 2 = AB 2 + BC 2 – 2 BC.BD.

AB 2 = AD 2 + DB 2
We can write it as;
⇒ AD 2 = AB 2 − DB 2 ……………….. (i)
By applying Pythagoras Theorem in ∆ADC, we get,
AD 2 + DC 2 = AC 2
From equation (i),
AB 2 − BD 2 + DC 2 = AC 2
AB 2 − BD 2 + (BC − BD) 2 = AC 2
AC 2 = AB 2 − BD 2 + BC 2 + BD 2 −2BC × BD
AC 2 = AB 2 + BC 2 − 2BC × BD
5. In Figure, AD is a median of a triangle ABC and AM ⊥ BC. Prove that :
(i) AC 2 = AD 2 + BC.DM + 2 (BC/2) 2
(ii) AB 2 = AD 2 – BC.DM + 2 (BC/2) 2
(iii) AC 2 + AB 2 = 2 AD 2 + ½ BC 2

(i) By applying Pythagoras Theorem in ∆AMD, we get,
AM 2 + MD 2 = AD 2 ………………. (i)
Again, by applying Pythagoras Theorem in ∆AMC, we get,
AM 2 + MC 2 = AC 2
AM 2 + (MD + DC) 2 = AC 2
(AM 2 + MD 2 ) + DC 2 + 2MD.DC = AC 2
From equation(i), we get,
AD 2 + DC 2 + 2MD.DC = AC 2
Since, DC=BC/2, thus, we get,
AD 2 + (BC/2) 2 + 2MD.(BC/2) 2 = AC 2
AD 2 + (BC/2) 2 + 2MD × BC = AC 2
(ii) By applying Pythagoras Theorem in ∆ABM, we get;
AB 2 = AM 2 + MB 2
= (AD 2 − DM 2 ) + MB 2
= (AD 2 − DM 2 ) + (BD − MD) 2
= AD 2 − DM 2 + BD 2 + MD 2 − 2BD × MD
= AD 2 + BD 2 − 2BD × MD
= AD 2 + (BC/2) 2 – 2(BC/2) MD
= AD 2 + (BC/2) 2 – BC MD
(iii) By applying Pythagoras Theorem in ∆ABM, we get,
AM 2 + MB 2 = AB 2 ………………….… (i)
By applying Pythagoras Theorem in ∆AMC, we get,
AM 2 + MC 2 = AC 2 …………………..… (ii)
Adding both the equations (i) and (ii), we get,
2AM 2 + MB 2 + MC 2 = AB 2 + AC 2
2AM 2 + (BD − DM) 2 + (MD + DC) 2 = AB 2 + AC 2
2AM 2 +BD 2 + DM 2 − 2BD.DM + MD 2 + DC 2 + 2MD.DC = AB 2 + AC 2
2AM 2 + 2MD 2 + BD 2 + DC 2 + 2MD (− BD + DC) = AB 2 + AC 2
2(AM 2 + MD 2 ) + (BC/2) 2 + (BC/2) 2 + 2MD (-BC/2 + BC/2) 2 = AB 2 + AC 2
2AD 2 + BC 2 /2 = AB 2 + AC 2
6. Prove that the sum of the squares of the diagonals of parallelogram is equal to the sum of the squares of its sides.
Let us consider, ABCD be a parallelogram. Now, draw perpendicular DE on extended side of AB, and draw a perpendicular AF meeting DC at point F.

By applying Pythagoras Theorem in ∆DEA, we get,
DE 2 + EA 2 = DA 2 ……………….… (i)
By applying Pythagoras Theorem in ∆DEB, we get,
DE 2 + EB 2 = DB 2
DE 2 + (EA + AB) 2 = DB 2
(DE 2 + EA 2 ) + AB 2 + 2EA × AB = DB 2
DA 2 + AB 2 + 2EA × AB = DB 2 ……………. (ii)
By applying Pythagoras Theorem in ∆ADF, we get,
AD 2 = AF 2 + FD 2
Again, applying Pythagoras theorem in ∆AFC, we get,
AC 2 = AF 2 + FC 2 = AF 2 + (DC − FD) 2
= AF 2 + DC 2 + FD 2 − 2DC × FD
= (AF 2 + FD 2 ) + DC 2 − 2DC × FD AC 2
AC 2 = AD 2 + DC 2 − 2DC × FD ………………… (iii)
Since ABCD is a parallelogram,
AB = CD ………………….…(iv)
And BC = AD ………………. (v)
In ∆DEA and ∆ADF,
∠DEA = ∠AFD (Each 90°)
∠EAD = ∠ADF (EA || DF)
AD = AD (Common Angles)
∴ ∆EAD ≅ ∆FDA (AAS congruence criterion)
⇒ EA = DF ……………… (vi)
Adding equations (i) and (iii), we get,
DA 2 + AB 2 + 2EA × AB + AD 2 + DC 2 − 2DC × FD = DB 2 + AC 2
DA 2 + AB 2 + AD 2 + DC 2 + 2EA × AB − 2DC × FD = DB 2 + AC 2
From equation (iv) and (vi),
BC 2 + AB 2 + AD 2 + DC 2 + 2EA × AB − 2AB × EA = DB 2 + AC 2
AB 2 + BC 2 + CD 2 + DA 2 = AC 2 + BD 2
7. In Figure, two chords AB and CD intersect each other at the point P. Prove that :
(i) ∆APC ~ ∆ DPB
(ii) AP . PB = CP . DP

Firstly, let us join CB, in the given figure.
(i) In ∆APC and ∆DPB,
∠APC = ∠DPB (Vertically opposite angles)
∠CAP = ∠BDP (Angles in the same segment for chord CB)
∆APC ∼ ∆DPB (AA similarity criterion)
(ii) In the above, we have proved that ∆APC ∼ ∆DPB
We know that the corresponding sides of similar triangles are proportional.
∴ AP/DP = PC/PB = CA/BD
⇒AP/DP = PC/PB
∴AP. PB = PC. DP
8. In Fig. 6.62, two chords AB and CD of a circle intersect each other at the point P (when produced) outside the circle. Prove that:
(i) ∆ PAC ~ ∆ PDB
(ii) PA . PB = PC . PD.

(i) In ∆PAC and ∆PDB,
∠P = ∠P (Common Angles)
As we know, exterior angle of a cyclic quadrilateral is ∠PCA and ∠PBD is opposite interior angle, which are both equal.
∠PAC = ∠PDB
Thus, ∆PAC ∼ ∆PDB(AA similarity criterion)
(ii) We have already proved above,
∆APC ∼ ∆DPB
AP/DP = PC/PB = CA/BD
AP/DP = PC/PB
∴ AP. PB = PC. DP
9. In Figure, D is a point on side BC of ∆ ABC such that BD/CD = AB/AC. Prove that AD is the bisector of ∠ BAC.

In the given figure, let us extend BA to P such that;
Now join PC.

Given, BD/CD = AB/AC
⇒ BD/CD = AP/AC
By using the converse of basic proportionality theorem, we get,
∠BAD = ∠APC (Corresponding angles) ……………….. (i)
And, ∠DAC = ∠ACP (Alternate interior angles) …….… (ii)
By the new figure, we have;
⇒ ∠APC = ∠ACP ……………………. (iii)
On comparing equations (i), (ii), and (iii), we get,
∠BAD = ∠APC
Therefore, AD is the bisector of the angle BAC.
10. Nazima is fly fishing in a stream. The tip of her fishing rod is 1.8 m above the surface of the water and the fly at the end of the string rests on the water 3.6 m away and 2.4 m from a point directly under the tip of the rod. Assuming that her string (from the tip of her rod to the fly) is taut, how much string does she have out (see Figure)? If she pulls in the string at the rate of 5 cm per second, what will be the horizontal distance of the fly from her after 12 seconds?

Let us consider, AB is the height of the tip of the fishing rod from the water surface and BC is the
horizontal distance of the fly from the tip of the fishing rod. Therefore, AC is now the length of the string.

To find AC, we have to use Pythagoras theorem in ∆ABC, is such way;
AC 2 = AB 2 + BC 2
AB 2 = (1.8 m) 2 + (2.4 m) 2
AB 2 = (3.24 + 5.76) m 2
AB 2 = 9.00 m 2
⟹ AB = √9 m = 3m
Thus, the length of the string out is 3 m.
As its given, she pulls the string at the rate of 5 cm per second.
Therefore, string pulled in 12 seconds = 12 × 5 = 60 cm = 0.6 m

Let us say now, the fly is at point D after 12 seconds.
Length of string out after 12 seconds is AD.
AD = AC − String pulled by Nazima in 12 seconds
= (3.00 − 0.6) m
In ∆ADB, by Pythagoras Theorem,
AB 2 + BD 2 = AD 2
(1.8 m) 2 + BD 2 = (2.4 m) 2
BD 2 = (5.76 − 3.24) m 2 = 2.52 m 2
BD = 1.587 m
Horizontal distance of fly = BD + 1.2 m
= (1.587 + 1.2) m = 2.787 m
NCERT Solutions for Class 10 Maths Chapter 6 – Triangles
NCERT Solutions Class 10 Maths Chapter 6 , Triangles, is part of the Unit Geometry, which constitutes 15 marks of the total marks of 80. On the basis of the updated CBSE Class 10 Syllabus for 2023-24, this chapter belongs to the Unit-Geometry and has the second-highest weightage. Hence, having a clear understanding of the concepts, theorems and problem-solving methods in this chapter is mandatory to score well in the board examination of Class 10 Maths.
Main topics covered in this chapter include:
6.1 introduction.
From your earlier classes, you are familiar with triangles and many of their properties. In Class 9, you have studied congruence of triangles in detail. In this chapter, we shall study about those figures which have the same shape, but not necessarily the same size. Two figures having the same shape (and not necessarily the same size) are called similar figures. In particular, we shall discuss the similarity of triangles and apply this knowledge in giving a simple proof of Pythagoras Theorem learnt earlier.
6.2 Similar Figures
In Class 9, you have seen that all circles with the same radii are congruent, all squares with the same side lengths are congruent and all equilateral triangles with the same side lengths are congruent. The topic explains similarity of figures by performing the relevant activity. Similar figures are two figures having the same shape, but not necessarily the same size.
6.3 Similarity of Triangles
The topic recalls triangles and its similarities. Two triangles are similar if (i) their corresponding angles are equal and (ii) their corresponding sides are in the same ratio (or proportion). It explains Basic Proportionality Theorem and different theorems are discussed performing various activities.
6.4 Criteria for Similarity of Triangles
In the previous section, we stated that two triangles are similar (i) their corresponding angles are equal and (ii) their corresponding sides are in the same ratio (or proportion). The topic discusses the criteria for similarity of triangles referring to the topics we have studied in earlier classes. It also contains different theorems explained with proper examples.
6.5 Areas of Similar Triangles
You have learnt that in two similar triangles, the ratio of their corresponding sides is the same. The topic Areas of Similar Triangles consists of theorem and relatable examples to prove the theorem.
6.6 Pythagoras Theorem
You are already familiar with the Pythagoras Theorem from your earlier classes. You have also seen proof of this theorem in Class 9. Now, we shall prove this theorem using the concept of similarity of triangles. Hence, the theorem is verified through some activities, and you can make use of it while solving certain problems.
6.7 Summary
The summary contains the points you have studied in the chapter. Going through the points mentioned in the summary will help you to recollect all the important concepts and theorems of the chapter.
List of Exercises in NCERT Class 10 Maths Chapter 6:
Exercise 6.1 Solutions 3 Questions (3 Short Answer Questions)
Exercise 6.2 Solutions 10 Questions (9 Short Answer Questions, 1 Long Answer Question)
Exercise 6.3 Solutions 16 Questions (1 main question with 6 sub-questions, 12 Short Answer Questions, 3 Long Answer Questions)
Exercise 6.4 Solutions 9 Questions (2 Short Answer with Reasoning Questions, 5 Short Answer Questions, 2 Long Answer Questions)
Exercise 6.5 Solutions 17 Questions (15 Short Answer Questions, 2 Long Answer Questions)
Exercise 6.6 Solutions 10 Questions (5 Short Answer Questions, 5 Long Answer Questions)
Triangle is one of the most interesting and exciting chapters of the unit Geometry as it takes us through the different aspects and concepts related to the geometrical figure triangle. A triangle is a plane figure that has three sides and three angles. This chapter covers various topics and sub-topics related to triangles, including a detailed explanation of similar figures, different theorems related to the similarities of triangles with proof, and the areas of similar triangles. The chapter concludes by explaining the Pythagoras theorem and the ways to use it in solving problems. Read and learn Chapter 6 of the Class 10 Maths NCERT textbook to learn more about Triangles and the concepts covered in it. Ensure to learn the NCERT Solutions for Class 10 effectively to score high in the board examinations.
Key Features of NCERT Solutions for Class 10 Maths Chapter 6 – Triangles
- Helps to ensure that the students use the concepts in solving the problems.
- Encourages the children to come up with diverse solutions to problems.
- Hints are given for those questions which are difficult to solve.
- Helps the students in checking if the solutions they gave for the questions are correct or not.
Disclaimer –
Dropped Topics –
Frequently Asked Questions on NCERT Solutions for Class 10 Maths Chapter 6
Why should we learn all the concepts present in ncert solutions for class 10 maths chapter 6, list out the important topics present in ncert solutions for class 10 maths chapter 6., how many exercises are there in ncert solutions for class 10 maths chapter 6, leave a comment cancel reply.
Your Mobile number and Email id will not be published. Required fields are marked *
Request OTP on Voice Call
Post My Comment
Register with BYJU'S & Download Free PDFs
Register with byju's & watch live videos.

Class 10 Maths Case Study Questions Chapter 6 Triangles PDF Download
- Post author: studyrate
- Post published:
- Post category: class 10th
- Post comments: 0 Comments
Case study Questions in the Class 10 Mathematics Chapter 6 are very important to solve for your exam. Class 10 Maths Chapter 6 Case Study Questions have been prepared for the latest exam pattern. You can check your knowledge by solving Class 10 Maths Case Study Questions Chapter 6 Triangles
Join our Telegram Channel, there you will get various e-books for CBSE 2024 Boards exams for Class 9th, 10th, 11th, and 12th.

In CBSE Class 10 Maths Paper, Students will have to answer some questions based on Assertion and Reason. There will be a few questions based on case studies and passage-based as well. In that, a paragraph will be given, and then the MCQ questions based on it will be asked.
Triangles Case Study Questions With Answers
Here, we have provided case-based/passage-based questions for Class 10 Maths Chapter 6 Triangles
Case Study/Passage-Based Questions
Case Study 1:
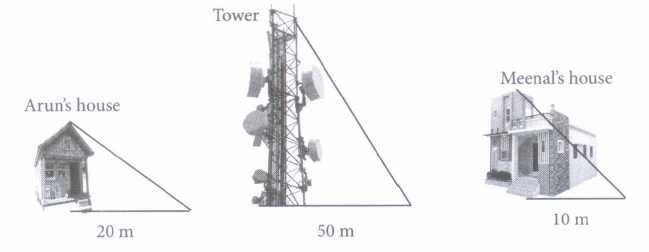
(i) What is the height of the tower?
| (a) 100 m | (b) 50 m | (c) 15 m | (d) 45 m |
Answer: (a) 100 m
(ii) What will be the length of the shadow of the tower when Meenal’s house casts a shadow of 15 m?
| (a) 45 m | (b) 70 m | (c) 75 m | (d) 72 m |
Answer: (c) 75 m
(iii) Height of Aruns house is
| (a) 80 m | (b) 75 m | (c) 60 m | (d) 40 m |
Answer: (d) 40 m
(iv) If the tower casts a shadow of 40 rn, then find the length of the shadow of Arun’s house
| (a) 18 m | (b) 17 m | (c) 16 m | (d) 14 m |
Answer: (b) 17 m
(v) If the tower casts a shadow of 40 m, then what will be the length of the shadow of Meenal’s house?
| (a) 7 m | (b) 9 m | (c) 4 m | (d) 8 m |
Answer: (d) 8 m
Case Study 2:
Rohan wants to measure the distance of a pond during the visit to his native. He marks points A and B on the opposite edges of a pond as shown in the figure below. To find the distance between the points, he makes a right-angled triangle using a rope connecting B with another point C is a distance of 12m, connecting C to point D at a distance of 40m from point C, and connecting D to the point A which is are a distance of 30m from D such the ∠ ADC=90 0 .

1. Which property of geometry will be used to find the distance AC?
a) Similarity of triangles
b) Thales Theorem
c) Pythagoras Theorem
d) Area of similar triangles
Answer: c)Pythagoras Theorem
2. What is the distance AC?
Answer: a) 50m
3. Which is the following does not form a Pythagoras triplet?
a) (7, 24, 25)
b) (15, 8, 17)
c) (5, 12, 13)
d) (21, 20, 28)
Answer: d) (21, 20, 28)
4. Find the length AB?
Answer: b) 38m
5. Find the length of the rope used.
Answer: c)82m
Hope the information shed above regarding Case Study and Passage Based Questions for Class 10 Maths Chapter 6 Triangles with Answers Pdf free download has been useful to an extent. If you have any other queries about CBSE Class 10 Maths Triangles Case Study and Passage-Based Questions with Answers, feel free to comment below so that we can revert back to us at the earliest possible By Team Study Rate
You Might Also Like
Class 10 english footprints without feet mcq questions with answers, extra questions of class 10 science chapter 11 human eye and colourful world pdf download, gk olympiad class 10 – syllabus, books, pyq, leave a reply cancel reply.
Save my name, email, and website in this browser for the next time I comment.
Class 10 Maths Chapter 6 Case Based Questions - Triangles
| 1 Crore+ students have signed up on EduRev. Have you? |
Case Study - 1
Case Study - 2
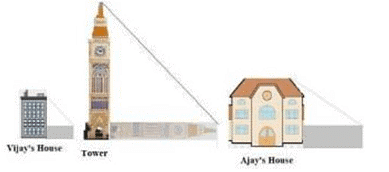
Vijay is trying to find the average height of a tower near his house. He is using the properties of similar triangles.The height of Vijay’s house if 20m when Vijay’s house casts a shadow 10m long on the ground. At the same time, the tower casts a shadow 50m long on the ground and the house of Ajay casts 20m shadow on the ground.
Q1: What is the height of the tower? (a) 20m (b) 50m (c) 100m (d) 200m Ans: (c) Explanation: The properties of similar triangles state that the ratios of the corresponding sides of the triangles are equal. Hence the ratio of the height of Vijay's house to the length of its shadow equals to the ratio of the height of the tower to the length of its shadow. Therefore, the height of the tower can be calculated as follows: Height of the tower = (Height of Vijay's house / Length of Vijay's house's shadow) * Length of the tower's shadow = (20m / 10m) * 50m = 100m Q2: What will be the length of the shadow of the tower when Vijay’s house casts a shadow of 12m? (a) 75m (b) 50m (c) 45m (d) 60m Ans: (d) Explanation: Applying the same method, we can find the length of the shadow of the tower. Length of the tower's shadow = (Length of Vijay's house's shadow / Height of Vijay's house) * Height of the tower = (12m / 20m) * 100m = 60m Q3: What is the height of Ajay’s house? (a) 30m (b) 40m (c) 50m (d) 20m Ans: (b) Explanation: Similarly, the height of Ajay's house can be calculated as follows: Height of Ajay's house = (Height of Vijay's house / Length of Vijay's house's shadow) * Length of Ajay's house's shadow = (20m / 10m) * 20m = 40m Q4: When the tower casts a shadow of 40m, same time what will be the length of the shadow of Ajay’s house? (a) 16m (b) 32m (c) 20m (d) 8m Ans: (a) Explanation: Length of Ajay's house's shadow = (Length of the tower's shadow / Height of the tower) * Height of Ajay's house = (40m / 100m) * 40m = 16m Q5: When the tower casts a shadow of 40m, same time what will be the length of the shadow of Vijay’s house? (a) 15m (b) 32m (c) 16m (d) 8m Ans: (d) Explanation: Length of Vijay's house's shadow = (Length of the tower's shadow / Height of the tower) * Height of Vijay's house = (40m / 100m) * 20m = 8m
Top Courses for Class 10
Viva Questions
Study material, practice quizzes, objective type questions, video lectures, extra questions, sample paper, important questions, semester notes, previous year questions with solutions, shortcuts and tricks, mock tests for examination, past year papers.

Case Based Questions: Triangles Free PDF Download
Importance of case based questions: triangles, case based questions: triangles notes, case based questions: triangles class 10, study case based questions: triangles on the app.
| cation olution |
| Join the 10M+ students on EduRev |
Welcome Back
Create your account for free.

Forgot Password
Unattempted tests, change country, practice & revise.
Not Able To Find Desired Paper or Worksheet SEARCH
Find papers & worksheets search, class 10 case study questions with answers - triangles.
- Mathematics

- (0) Comments
- 15 Downloads
Related Papers
Click to view more related papers, display_name = "class 11" && $paper->display_name = "class 12") { // echo $paper->display_name." questions papers and worksheets"; } //else { // echo $paper->display_name." sample papers and previous year papers"; //} //>, cbse worksheets for class 10 maths.
Download free printable worksheets for CBSE Class 10 Maths with important chapter wise questions as per Latest NCERT Syllabus. These Worksheets help Grade 10 students practice Maths Important Questions and exercises on various topics like Coordinate Geometry, Probability, Quadratic Equations, Statistics, Surface Area, Arithmetic Progression, Polynomials, Linear Equation in two Variables, Real Numbers. These free PDF download of Class 10 Maths worksheets consist of visual simulations to help your child visualize concepts being taught and reinforce their learning.
Get free Kendriya Vidyalaya Class 10 Maths Worksheets shared by teachers, parents & students to understand the concepts. All the necessary topics are covered in these 10th grade worksheets. These class 10 Maths worksheets provide skills and experience necessary to ace in Exams
Total Papers :
| Class 10 Maths Marks Distribution | |
|---|---|
| Units | Marks |
| Number Systems | 06 |
| Algebra | 20 |
| Coordinate Geometry | 06 |
| Geometry | 15 |
| Trigonometry | 12 |
| Mensuration | 10 |
| Statistics & Probability | 11 |
| Internal Assessment | 20 |
| Total | 100 |
CBSE Class 10 Maths Syllabus
- Real Numbers
- Polynomials
- Pair of Linear Equations in Two Variables
- Quadratic Equations
- Arithmetic Progressions
- Coordinate Geometry
- Probability
- Introduction to Trigonometry
- Some Applications of Trigonometry
- Constructions
- Area Related to Circles
- Surface Areas and Volumes
UNIT I: NUMBER SYSTEMS
1. REAL NUMBER
Fundamental Theorem of Arithmetic - statements after reviewing work done earlier and after illustrating and motivating through examples, Proofs of irrationality.
UNIT II: ALGEBRA
- POLYNOMIALS Zeros of a polynomial. Relationship between zeros and coefficients of quadratic polynomials.
- PAIR OF LINEAR EQUATIONS IN TWO VARIABLES Pair of linear equations in two variables and graphical method of their solution, consistency/inconsistency. Algebraic conditions for number of solutions. Solution of a pair of linear equations in two variables algebraically - by substitution, by elimination. Simple situational problems.
- QUADRATIC EQUATIONS Standard form of a quadratic equation ax2 + bx + c = 0, (a ≠ 0). Solutions of quadratic equations (only real roots) by factorization, and by using quadratic formula. Relationship between discriminant and nature of roots. Situational problems based on quadratic equations related to day to day activities to be incorporated.
- ARITHMETIC PROGRESSIONS Motivation for studying Arithmetic Progression Derivation of the nth term and sum of the first n terms of A.P. and their application in solving daily life problems.
UNIT III: COORDINATE GEOMETRY Coordinate Geometry Review: Concepts of coordinate geometry, graphs of linear equations. Distance formula. Section formula (internal division).
UNIT IV: GEOMETRY
- TRIANGLES Definitions, examples, counter examples of similar triangles. 1. (Prove) If a line is drawn parallel to one side of a triangle to intersect the other two sides in distinct points, the other two sides are divided in the same ratio. 2. (Motivate) If a line divides two sides of a triangle in the same ratio, the line is parallel to the third side. 3.(Motivate) If in two triangles, the corresponding angles are equal, their corresponding sides are proportional and the triangles are similar. 4.(Motivate) If the corresponding sides of two triangles are proportional, their corresponding angles are equal and the two triangles are similar. 5.(Motivate) If one angle of a triangle is equal to one angle of another triangle and the sides including these angles are proportional, the two triangles are similar.
- CIRCLES Tangent to a circle at, point of contact 1. (Prove) The tangent at any point of a circle is perpendicular to the radius through the point of contact. 2.(Prove) The lengths of tangents drawn from an external point to a circle are equal.
UNIT V: TRIGONOMETRY
- INTRODUCTION TO TRIGONOMETRY Trigonometric ratios of an acute angle of a right-angled triangle. Proof of their existence (well defined); motivate the ratios whichever are defined at 0° and 90°. Values of the trigonometric ratios of 30°, 45°, and 60°. Relationships between the ratios.
- TRIGONOMETRIC IDENTITIES Proof and applications of the identity sin2A + cos2A = 1 . Only simple identities to be given.
- HEIGHTS AND DISTANCES: Angle of elevation, Angle of Depression. (10)Periods Simple problems on heights and distances. Problems should not involve more than two right triangles. Angles of elevation / depression should be only 30°, 45°, and 60°.
UNIT VI: MENSURATION
- AREAS RELATED TO CIRCLES Area of sectors and segments of a circle. Problems based on areas and perimeter / circumference of the above said plane figures. (In calculating area of segment of a circle, problems should be restricted to central angle of 60°, 90° and 120° only.
- SURFACE AREAS AND VOLUMES Surface areas and volumes of combinations of any two of the following: cubes, cuboids, spheres, hemispheres and right circular cylinders/cones.
UNIT VII: STATISTICS AND PROBABILITY
- STATISTICS Mean, median and mode of grouped data (bimodal situation to be avoided).
- PROBABILITY (10) Periods Classical definition of probability. Simple problems on finding the probability of an event.
PRESCRIBED BOOKS:
- Mathematics - Textbook for class IX - NCERT Publication
- Mathematics - Textbook for class X - NCERT Publication
- Guidelines for Mathematics Laboratory in Schools, class IX - CBSE Publication
- Guidelines for Mathematics Laboratory in Schools, class X - CBSE Publication
- Laboratory Manual - Mathematics, secondary stage - NCERT Publication
- Mathematics exemplar problems for class IX, NCERT publication.
- Mathematics exemplar problems for class X, NCERT publication.
Structure of CBSE Maths Sample Paper for Class 10 is
| Type of Question | Marks per Question | Total No. of Questions | Total Marks |
|---|---|---|---|
| Objective Type Questions | 1 | 20 | 20 |
| Short Answer Type Questions - I | 2 | 6 | 12 |
| Short Answer Type Questions - II | 3 | 8 | 24 |
| Long Answer Type Questions | 4 | 6 | 24 |
| Total | 40 | 80 | |
For Preparation of board exams students can also check out other resource material
CBSE Class 10 Maths Sample Papers
Important Questions for Class 10 Maths Chapter Wise
Maths Revision Notes for class 10
Previous Year Question Paper CBSE Class 10 Maths
Worksheets of Other Subjects of Class 10
Why do one Children need Worksheets for Practice ?
It is very old saying that one can build a large building if the foundation is strong and sturdy. This holds true for studies also. Worksheets are essential and help students in the in-depth understanding of fundamental concepts. Practicing solving a lot of worksheets, solving numerous types of questions on each topic holds the key for success. Once basic concepts and fundamentals have been learnt, the next thing is to learn their applications by practicing problems. Practicing the problems helps us immensely to gauge how well we have understood the concepts.
There are times when students just run through any particular topic with casual awareness there by missing out on a few imperative “between the lines” concepts. Such things are the major causes of weak fundamental understandings of students. So in such cases Worksheets act as a boon and critical helpful tool which gauges the in-depth understanding of children highlighting doubts and misconceptions, if any.
Worksheets classifies the important aspects of any topic or chapter taught in the class in a very easy manner and increases the awareness amongst students.When students try to solve a worksheet they get to understand what are the key important factors which needs the main focus.Sometimes it happens that due to shortage of time all the major points of any particular topic gets skipped in the class or teacher rushes through , due to shortage of time. A worksheet thus provides a framework for the entire chapter and can help covering those important aspects which were rushed in the class and ensure that students record and understand all key items.
In a class of its say 40 students howsoever teacher tries to be active and work towards making each student understand whatever she has to teach in the class but there are always some students who tend to be in their own world and they wander in their thoughts.Worksheets which are provided timely to all the students, causes them to focus on the material at hand. it’s simply the difference between passive and active learning. Worksheets of this type can be used to introduce new material, particularly material with many new definitions and terms.
Worksheets help students be focussed and attentive in the class because they know after the class is over they will be assigned a worksheet which they need to solve so if they miss or skip any point in the class they may not be able to solve the worksheet completely and thereby lose reputation in the class.
Often students revise the chapter at home reading their respective textbooks. Thus more often than not they do miss many important points. Worksheets thus can be used intentionally to help guide student’s to consult textbooks. Having students write out responses encourages their engagement with the textbooks, the questions chosen indicate areas on which to focus. Explicitly discussing the worksheets and why particular questions are asked helps students reflect on what is important.
Worksheets of Other Classes
To Prepare better for CBSE paperclass; ?> " title="Download Free CBSE Papers">Ribblu.com brings to you all the previous years papers & worksheets of subject; ?//> for CBSE paperclass; ?>. This CBSE paper and worksheet can be instrumental in students achieving maximum marks in their exams. These Papers and worksheets help students gain confidence and make them ready to face their school examinations. These Papers and worksheets school wise, covers important concepts from an examination perspective. Students and parents can download all the available papers & worksheets directly in the form of PDF. One can use these papers and worksheets to get extensive practice and familiarise themselves with the format of the question paper.
You can help other users
Be the first to write comment .
Upload papers and the more your paper get downloaded the more you earn the points
You may send papers on email [email protected] along with userid
- Downloaded by: Ranjana Rathaur
- Downloaded by: Neha Chandel
- Downloaded by: Gayathri Srinivasan
Rules and regulations for uploads
| 1. | The uploaded material should be original and not duplicated. |
| 2. | It should be clear, legible and genuine. The file type should be pdf for multiple pages and jpg is allowed only for single page document. Use apps like “cam scanner” for mobile capture, crop and edit photos and save it as pdf. One file should be complete in all aspects like one full question paper, Notes of a complete topic. File name should be self explanatory (eg. CBSE 10th 2012 hindi question paper by – ‘Name of the uploader’) |
| 3. | No copyright violations allowed. |
| 4. | Points and coupons will be given at the sole discretion of Ribblu. |
| 5. | Ribblu admin has the power to reject, remove, alter, approve, accept any material that is uploaded by the user without consent of owner. |
Write your comment
Report this paper, how to earn points.
Upload Papers / Worksheets and Earn 50 Points.
The uploaded material should be original paper or worksheet of any school. Check out some videos on how to upload papers on ribblu
Rate & Review your school and Earn 25 Points.
Review any school that you may be knowing and once your review is approved, you will be credited with 25 points.
Answer on question posted on JustAsk and earn 15 points.
JustAsk is a platform where you can help others to find answers of any question. Share your Knowledge. Answer questions and once approved you will earn 15 points
Complete your profile and earn upto 25 Points.
Edit and complete your user profile and earn points. The more details you submit, the more points you will earn.
Download Ribblu Mobile App and you will (Earn 20 Points) (one time only)
CBSE Schools
- CBSE Schools In Delhi
- CBSE Schools In Noida
- CBSE Schools In Greater Noida
- CBSE Schools In Faridabad
- CBSE Schools In Ghaziabad
- CBSE Schools In Gurgaon
- CBSE Schools In Mumbai
- CBSE Schools In Pune
- CBSE Schools In Bangalore
- CBSE Schools In Hyderabad
- CBSE Schools In Kolkata
- CBSE Schools In Chennai
- CBSE Schools In Patna
- CBSE Schools In Meerut
- CBSE Schools In Kanpur
- CBSE Schools In Indore
- CBSE Schools In Ludhiana
- CBSE Schools In Dehradun
Top Schools
- Schools In Delhi
- Schools In Noida
- Schools In Greater Noida
- Schools In Faridabad
- Schools In Ghaziabad
- Schools In Gurgaon
- Schools In Mumbai
- Schools In Pune
- Schools In Bangalore
- Schools In Hyderabad
- Schools In Kolkata
- Schools In Chennai
- Schools In Patna
- Schools In Meerut
- Schools In Kanpur
- Schools In Indore
- Schools In Ludhiana
- Schools In Dehradun
Other Schools
- Pre Nursery Schools In Noida
- Day Boarding Schools In Noida
- Pre Nursery Schools In Gurgaon
- Pre Nursery Schools In Delhi
- Play Schools In Delhi
- Day Boarding Schools In Delhi
CBSE Papers
- CBSE Class 1 Sample Papers
- CBSE Class 2 Sample Papers
- CBSE Class 3 Sample Papers
- CBSE Class 4 Sample Papers
- CBSE Class 5 Sample Papers
- CBSE Class 6 Sample Papers
- CBSE Class 7 Sample Papers
- CBSE Class 8 Sample Papers
Paper Categories
- Question Bank
- Question Papers
- Revision Notes
- Sample Papers
- Test Papers
- CBSE Class 9 Sample Papers
- CBSE Class 10 Sample Papers
- CBSE Class 11 Sample Papers
- CBSE Class 12 Sample Papers


Gurukul of Excellence
Classes for Physics, Chemistry and Mathematics by IITians
Join our Telegram Channel for Free PDF Download
Case Study Questions for Class 10 Maths Chapter 6 Triangles
- Last modified on: 12 months ago
- Reading Time: 4 Minutes
Case Study Questions:
Question 1:
Vijay is trying to find the average height of a tower near his house. He is using the properties of similar triangles. The height of Vijay’s house if 20 m when Vijay’s house casts a shadow 10 m long on the ground. At the same time, the tower casts a shadow 50m long on the ground and the house of Ajay casts 20 m shadow on the ground.

(i) What is the height of the tower? (a) 20 m (b) 50 m (c) 100 m (d) 200 m
(ii) What will be the length of the shadow of the tower when Vijay’s house casts a shadow of 12m? (a) 75 m (b) 50 m (c) 45 m (d) 60 m
(iii) What is the height of Ajay’s house? (a) 30 m (b) 40 m (c) 50 m (d) 20 m
(iv) When the tower casts a shadow of 40m, same time what will be the length of the shadow of Ajay’s house? (a) 16 m (b) 32 m (c) 20 m (d) 8 m
(v) When the tower casts a shadow of 40m, same time what will be the length of the shadow of Vijay’s house? (a) 15 m (b) 32 m (c) 16 m (d) 8 m
✨ Free Quizzes, Test Series and Learning Videos for CBSE Class 10 Maths
You may also like:
Chapter 1 Real Numbers Chapter 2 Polynomials Chapter 3 Pair of Linear Equations in Two Variables C hapter 4 Quadratic Equations Chapter 5 Arithmetic Progressions Chapter 6 Triangles Chapter 7 Coordinate Geometry Chapter 8 Introduction to Trigonometry Chapter 9 Some Applications of Trigonometry Chapter 10 Circles Chapter 11 Constructions Chapter 12 Areas Related to Circles Chapter 13 Surface Areas and Volumes Chapter 14 Statistics Chapter 15 Probability
Download CBSE Books
Exam Special Series:
- Sample Question Paper for CBSE Class 10 Science (for 2024)
- Sample Question Paper for CBSE Class 10 Maths (for 2024)
- CBSE Most Repeated Questions for Class 10 Science Board Exams
- CBSE Important Diagram Based Questions Class 10 Physics Board Exams
- CBSE Important Numericals Class 10 Physics Board Exams
- CBSE Practical Based Questions for Class 10 Science Board Exams
- CBSE Important “Differentiate Between” Based Questions Class 10 Social Science
- Sample Question Papers for CBSE Class 12 Physics (for 2024)
- Sample Question Papers for CBSE Class 12 Chemistry (for 2024)
- Sample Question Papers for CBSE Class 12 Maths (for 2024)
- Sample Question Papers for CBSE Class 12 Biology (for 2024)
- CBSE Important Diagrams & Graphs Asked in Board Exams Class 12 Physics
- Master Organic Conversions CBSE Class 12 Chemistry Board Exams
- CBSE Important Numericals Class 12 Physics Board Exams
- CBSE Important Definitions Class 12 Physics Board Exams
- CBSE Important Laws & Principles Class 12 Physics Board Exams
- 10 Years CBSE Class 12 Chemistry Previous Year-Wise Solved Papers (2023-2024)
- 10 Years CBSE Class 12 Physics Previous Year-Wise Solved Papers (2023-2024)
- 10 Years CBSE Class 12 Maths Previous Year-Wise Solved Papers (2023-2024)
- 10 Years CBSE Class 12 Biology Previous Year-Wise Solved Papers (2023-2024)
- ICSE Important Numericals Class 10 Physics BOARD Exams (215 Numericals)
- ICSE Important Figure Based Questions Class 10 Physics BOARD Exams (230 Questions)
- ICSE Mole Concept and Stoichiometry Numericals Class 10 Chemistry (65 Numericals)
- ICSE Reasoning Based Questions Class 10 Chemistry BOARD Exams (150 Qs)
- ICSE Important Functions and Locations Based Questions Class 10 Biology
- ICSE Reasoning Based Questions Class 10 Biology BOARD Exams (100 Qs)
✨ Join our Online JEE Test Series for 499/- Only (Web + App) for 1 Year
✨ Join our Online NEET Test Series for 499/- Only for 1 Year
Leave a Reply Cancel reply
Join our Online Test Series for CBSE, ICSE, JEE, NEET and Other Exams

Editable Study Materials for Your Institute - CBSE, ICSE, State Boards (Maharashtra & Karnataka), JEE, NEET, FOUNDATION, OLYMPIADS, PPTs
Discover more from Gurukul of Excellence
Subscribe now to keep reading and get access to the full archive.
Type your email…
Continue reading

- Andhra Pradesh
- Chhattisgarh
- West Bengal
- Madhya Pradesh
- Maharashtra
- Jammu & Kashmir
- NCERT Books 2022-23
- NCERT Solutions
- NCERT Notes
- NCERT Exemplar Books
- NCERT Exemplar Solution
- States UT Book
- School Kits & Lab Manual
- NCERT Books 2021-22
- NCERT Books 2020-21
- NCERT Book 2019-2020
- NCERT Book 2015-2016
- RD Sharma Solution
- TS Grewal Solution
- TR Jain Solution
- Selina Solution
- Frank Solution
- Lakhmir Singh and Manjit Kaur Solution
- I.E.Irodov solutions
- ICSE - Goyal Brothers Park
- ICSE - Dorothy M. Noronhe
- Micheal Vaz Solution
- S.S. Krotov Solution
- Evergreen Science
- KC Sinha Solution
- ICSE - ISC Jayanti Sengupta, Oxford
- ICSE Focus on History
- ICSE GeoGraphy Voyage
- ICSE Hindi Solution
- ICSE Treasure Trove Solution
- Thomas & Finney Solution
- SL Loney Solution
- SB Mathur Solution
- P Bahadur Solution
- Narendra Awasthi Solution
- MS Chauhan Solution
- LA Sena Solution
- Integral Calculus Amit Agarwal Solution
- IA Maron Solution
- Hall & Knight Solution
- Errorless Solution
- Pradeep's KL Gogia Solution
- OP Tandon Solutions
- Sample Papers
- Previous Year Question Paper
- Important Question
- Value Based Questions
- CBSE Syllabus
- CBSE MCQs PDF
- Assertion & Reason
- New Revision Notes
- Revision Notes
- Question Bank
- Marks Wise Question
- Toppers Answer Sheets
- Exam Paper Aalysis
- Concept Map
- CBSE Text Book
- Additional Practice Questions
- Vocational Book
- CBSE - Concept
- KVS NCERT CBSE Worksheets
- Formula Class Wise
- Formula Chapter Wise
- JEE Previous Year Paper
- JEE Mock Test
- JEE Crash Course
- JEE Sample Papers
- Important Info
- SRM-JEEE Previous Year Paper
- SRM-JEEE Mock Test
- VITEEE Previous Year Paper
- VITEEE Mock Test
- BITSAT Previous Year Paper
- BITSAT Mock Test
- Manipal Previous Year Paper
- Manipal Engineering Mock Test
- AP EAMCET Previous Year Paper
- AP EAMCET Mock Test
- COMEDK Previous Year Paper
- COMEDK Mock Test
- GUJCET Previous Year Paper
- GUJCET Mock Test
- KCET Previous Year Paper
- KCET Mock Test
- KEAM Previous Year Paper
- KEAM Mock Test
- MHT CET Previous Year Paper
- MHT CET Mock Test
- TS EAMCET Previous Year Paper
- TS EAMCET Mock Test
- WBJEE Previous Year Paper
- WBJEE Mock Test
- AMU Previous Year Paper
- AMU Mock Test
- CUSAT Previous Year Paper
- CUSAT Mock Test
- AEEE Previous Year Paper
- AEEE Mock Test
- UPSEE Previous Year Paper
- UPSEE Mock Test
- CGPET Previous Year Paper
- Crash Course
- Previous Year Paper
- NCERT Based Short Notes
- NCERT Based Tests
- NEET Sample Paper
- Previous Year Papers
- Quantitative Aptitude
- Numerical Aptitude Data Interpretation
- General Knowledge
- Mathematics
- Agriculture
- Accountancy
- Business Studies
- Political science
- Enviromental Studies
- Mass Media Communication
- Teaching Aptitude
- Verbal Ability & Reading Comprehension
- Logical Reasoning & Data Interpretation
- NAVODAYA VIDYALAYA
- SAINIK SCHOOL (AISSEE)
- Mechanical Engineering
- Electrical Engineering
- Electronics & Communication Engineering
- Civil Engineering
- Computer Science Engineering
- CBSE Board News
- Scholarship Olympiad
- School Admissions
- Entrance Exams
- All Board Updates
- Miscellaneous
- State Wise Books
- Engineering Exam
- STATE WISE BOOKS
- ENGINEERING EXAM
- SCHOLARSHIP OLYMPIAD
- STATE BOOKS
CBSE Class 10 Maths Case Study
CBSE Board has introduced the case study questions for the ongoing academic session 2021-22. The board will ask the paper on the basis of a different exam pattern which has been introduced this year where 50% syllabus is occupied for MCQ for Term 1 exam. Selfstudys has provided below the chapter-wise questions for CBSE Class 10 Maths. Students must solve these case study based problems as soon as they are done with their syllabus.
These case studies are in the form of Multiple Choice Questions where students need to answer them as asked in the exam. The MCQs are not that difficult but having a deep and thorough understanding of NCERT Maths textbooks are required to answer these. Furthermore, we have provided the PDF File of CBSE Class 10 maths case study 2021-2022.
Class 10 Maths (Formula, Case Based, MCQ, Assertion Reason Question with Solutions)
In order to score good marks in the term 1 exam students must be aware of the Important formulas, Case Based Questions, MCQ and Assertion Reasons with solutions. Solving these types of questions is important because the board will ask them in the Term 1 exam as per the changed exam pattern of CBSE Class 10th.
Important formulas should be necessarily learned by the students because the case studies are solved with the help of important formulas. Apart from that there are assertion reason based questions that are important too.
| Real Number | |
| Polynomials ( ) | |
| Pair of Linear Equations in Two Variables (MCQ, Case-Based, Assertion & Reasoning) | |
| Coordinate Geometry (MCQ, Case-Based, Assertion & Reasoning) | |
| Triangles | |
| Introduction to Trigonometry (MCQ, Case-Based, Assertion & Reasoning) | |
| Areas Related to Circles (MCQ, Case-Based, Assertion & Reasoning) | |
| Probability (MCQ, Case-Based, Assertion & Reasoning) | |
| Quadratic Equation (MCQ) | |
| Arithmetic Progression (MCQ) | |
| Some Application of Trigonometry (MCQ) | |
| Circles (MCQ) | |
| Constructions (MCQ) | |
| Surface Areas and Volumes (MCQ) | |
| Statistics (MCQ) |
Assertion Reasoning is a kind of question in which one statement (Assertion) is given and its reason is given (Explanation of statement). Students need to decide whether both the statement and reason are correct or not. If both are correct then they have to decide whether the given reason supports the statement or not. In such ways, assertion reasoning questions are being solved. However, for doing so and getting rid of confusions while solving. Students are advised to practice these as much as possible.
For doing so we have given the PDF that has a bunch of MCQs questions based on case based, assertion, important formulas, etc. All the Multiple Choice problems are given with detailed explanations.
CBSE Class 10th Case study Questions
Recently CBSE Board has the exam pattern and included case study questions to make the final paper a little easier. However, Many students are nervous after hearing about the case based questions. They should not be nervous because case study are easy and given in the board papers to ease the Class 10th board exam papers. However to answer them a thorough understanding of the basic concepts are important. For which students can refer to the NCERT textbook.
Basically, case study are the types of questions which are developed from the given data. In these types of problems, a paragraph or passage is given followed by the 5 questions that are given to answer . These types of problems are generally easy to answer because the data are given in the passage and students have to just analyse and find those data to answer the questions.
CBSE Class 10th Assertion Reasoning Questions
These types of questions are solved by reading the statement, and given reason. Sometimes these types of problems can make students confused. To understand the assertion and reason, students need to know that there will be one statement that is known as assertion and another one will be the reason, which is supposed to be the reason for the given statement. However, it is students duty to determine whether the statement and reason are correct or not. If both are correct then it becomes important to check, does reason support the statement?
Moreover, to solve the problem they need to look at the given options and then answer them.
CBSE Class 10 Maths Case Based MCQ
CBSE Class 10 Maths Case Based MCQ are either Multiple Choice Questions or assertion reasons. To solve such types of problems it is ideal to use elimination methods. Doing so will save time and answering the questions will be much easier. Students preparing for the board exams should definitely solve these types of problems on a daily basis.
Also, the CBSE Class 10 Maths MCQ Based Questions are provided to us to download in PDF file format. All are developed as per the latest syllabus of CBSE Class Xth.
Class 10th Mathematics Multiple Choice Questions
Class 10 Mathematics Multiple Choice Questions for all the chapters helps students to quickly revise their learnings, and complete their syllabus multiple times. MCQs are in the form of objective types of questions whose 4 different options are given and one of them is a true answer to that problem. Such types of problems also aid in self assessment.
Case Study Based Questions of class 10th Maths are in the form of passage. In these types of questions the paragraphs are given and students need to find out the given data from the paragraph to answer the questions. The problems are generally in Multiple Choice Questions.
The Best Class 10 Maths Case Study Questions are available on Selfstudys.com. Click here to download for free.
To solve Class 10 Maths Case Studies Questions you need to read the passage and questions very carefully. Once you are done with reading you can begin to solve the questions one by one. While solving the problems you have to look at the data and clues mentioned in the passage.
In Class 10 Mathematics the assertion and reasoning questions are a kind of Multiple Choice Questions where a statement is given and a reason is given for that individual statement. Now, to answer the questions you need to verify the statement (assertion) and reason too. If both are true then the last step is to see whether the given reason support=rts the statement or not.

CBSE Class 10 Results 2024 : CBSE Class 10 Answer Book Photocopy Applications Open

CBSE 10th 2024-25 : Social Science Official Competency Focused Practice Questions released by CBSE


CBSE Class 10 Revaluation Application 2024 Process Begins : How to Apply, Fees, Direct Link & Step-by-Step Guide

CBSE Class 10 Result 2024 Latest Update : Verification of Marks, Revaluation & Photocopy of Answer Sheet; Check Complete Process

CBSE Class 10 Result 2024 Out: 93.60% Pass Percentage, JNV and Kendriya Vidyalaya shine with 99.09% score

CBSE Class 10th Result 2024 Out : Supplementary, Re-verification & Re-evaluation Date Released; Check Details Here

- NCERT Solutions for Class 12 Maths
- NCERT Solutions for Class 10 Maths
- CBSE Syllabus 2023-24
- Social Media Channels
- Login Customize Your Notification Preferences

- Second click on the toggle icon

Provide prime members with unlimited access to all study materials in PDF format.
Allow prime members to attempt MCQ tests multiple times to enhance their learning and understanding.
Provide prime users with access to exclusive PDF study materials that are not available to regular users.

NCERT Solutions for Class 6, 7, 8, 9, 10, 11 and 12
NCERT Solutions For Class 10 Maths Chapter 6 Triangles Ex 6.1
Get Free NCERT Solutions for Class 10 Maths Chapter 6 Ex 6.1 PDF. Triangles Class 10 Maths NCERT Solutions are extremely helpful while doing your homework. Exercise 6.1 Class 10 Maths NCERT Solutions were prepared by Experienced LearnCBSE.in Teachers. Detailed answers of all the questions in Chapter 6 Maths Class 10 Triangles Exercise 6.1 provided in NCERT TextBook.
You can also download Maths Class 10 to help you to revise complete syllabus and score more marks in your examinations.
Free download NCERT Solutions for Class 10 Maths Chapter 6 Exercise 6.1 Triangles PDF in Hindi Medium as well as in English Medium for CBSE, Uttarakhand, Bihar, MP Board, Gujarat Board, AP SSC, TS SSC and UP Board students, who are using NCERT Books based on updated CBSE Syllabus for the session 2019-20.
- Triangles Class 10 Mind Map
- Triangles Class 10 Ex 6.1
- Triangles Class 10 Ex 6.1 in Hindi Medium
Triangles Class 10 Ex 6.2
- Triangles Class 10 Ex 6.2 in Hindi Medium
Triangles Class 10 Ex 6.3
- Triangles Class 10 Ex 6.3 in Hindi Medium
Triangles Class 10 Ex 6.4
- Triangles Class 10 Ex 6.4 in Hindi Medium
Triangles Class 10 Ex 6.5
- Triangles Class 10 Ex 6.5 in Hindi Medium
Triangles Class 10 Ex 6.6
- Triangles Class 10 Ex 6.6 in Hindi Medium
- Extra Questions for Class 10 Maths Triangles
- Triangles Class 10 Notes Maths Chapter 6
- NCERT Exemplar Class 10 Maths Chapter 6 Triangles
- Important Questions for Class 10 Maths Chapter 6 Triangles
NCERT Solutions for Class 10 Maths Chapter 6 Triangles Ex Ex 6.1 are part of Class 10 Maths NCERT Solutions . Here we have given NCERT Solutions for Class 10 Maths Chapter 6 Triangles Exercise 6.1
| CBSE | |
| NCERT | |
| Class 10 | |
| Maths | |
| Chapter 6 | |
| Triangles | |
| Ex 6.1 | |
| 3 | |

NCERT Solutions For Class 10 Maths Chapter 6 Triangles Ex 6.1 in Hindi Medium
प्र. 1. कोष्ठकों में दिए शब्दों में से सही शब्दों का प्रयोग करते हुए, रिक्त स्थानों को भरिए : (i) सभी वृत्त …….. होते है| (सर्वांगसम, समरूप) (ii) सभी वर्ग…… होते हैं| (समरूप, सर्वांगसम) (iii) सभी …….. त्रिभुज समरूप होते है | (समद्विबाहु, समबाहु) (iv) भुजाओं की समान संख्या वाले दो बहुभुज समरूप होते हैं, यदि (i) उनके संगत कोण ……..हो तथा (ii) उनकी संगत ……भुजाएँ हों| (बराबर, समानुपाती| हलः (i) सभी वृत्त समरूप होते हैं। (ii) सभी वर्ग समरूप होते हैं। (iii) सभी समबाहु त्रिभुज समरूप होते हैं। (iv) भुजाओं की समान संख्या वाले दो बहुभुजे समरूप होते हैं, यदि (i) उनके संगत कोण बराबर हों तथा (ii) उनकी संगत समानुपाती भुजाएँ हों।
प्र० 2. निम्नलिखित युग्मों के दो भिन्न-भिन्न उदाहरण दीजिएः (i) समरूप आकृतियाँ (ii) ऐसी आकृतियाँ जो समरूप नहीं हैं। हलः (i) (a) दो वृत्त परस्पर समरूप होते हैं। (b) दो वर्ग परस्पर समरूप होते हैं। (ii) (a) एक वृत्त और एक त्रिभुज समरूप नहीं होते हैं। (b) एक समद्विबाहु त्रिभुज और एक विषमबाहु। त्रिभुज समरूप आकृतियाँ नहीं होती हैं।
प्र० 3. बताइए कि निम्न चतुर्भुज समरूप हैं या नहीं:
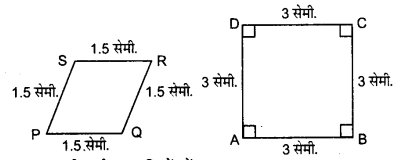
संगत भुजाएँ समानुपाती हैं, परन्तु इनके संगत कोण समान नहीं हैं। ये आकृतियाँ समरूप नहीं हैं।
Class 10 Maths Triangles Mind Map
Similar figures.
Two figures having the same shape but not necessarily the same size are called similar figures Two figures having the same shape as well as same size are called congruent figures Note that all congruent figures are similar but the similar figures need not be congruent.
Similarity of Polygons
Two polygons of the same number of sides are similar if (i) their corresponding angles are equal and (ii) their corresponding sides are in the same ratio (or proportion)
Similarity of Triangles
Two triangles are similar if (i) their corresponding angles are equal and (ii) their corresponding sides are in the same ratio (or proportion) Note : If the corresponding angles of two triangles are equal, then they are known as equiangular triangles. The ratio of any two corresponding sides in two equiangular triangles is always the same.
Basic Proportionality Theorem (BPT) and its Converse
Criteria For Similarity of Triangles
(i) AAA Similarity Criterion : If in two triangles, corresponding angles are equal then their corresponding sides are in the same ratio and hence the two triangles are similar. (ii) AA Similarity Criterion : If in two triangles, two angles of one triangle are respectively equal to the two angles of the other triangle, then the two triangles are similar. (iii) SSS Similarity Criterion : If in two triangles, corresponding sides are in the same ratio then their corresponding angles are equal and hence the triangles are similar. (iv) SAS Similarity Criterion : If one angle of a triangle is equal to one angle of another triangle and the sides including these angles are in the same ratio (proportion), then the two triangles are similar.
Areas of Similar Triangles
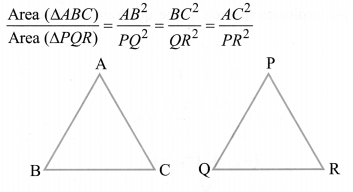
Pythagoras Theorem and its Converse
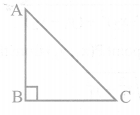
You can also download the free PDF of Class 10 Triangles NCERT Solutions or save the solution images and take the print out to keep it handy for your exam preparation.
Topics and Sub Topics in Class 10 Maths Chapter 6 Triangles:
| 6 | Triangles |
| 6.1 | Introduction |
| 6.2 | Similar Figures |
| 6.3 | Similarity Of Triangles |
| 6.4 | Criteria For Similarity Of Triangles |
| 6.5 | Areas Of Similar Triangles |
| 6.6 | Pythagoras Theorem |
| 6.7 | Summary |
More Resources for CBSE Class 10
NCERT Solutions
Ncert solutions for class 10 maths.
- NCERT Solutions for Class 10 Science
- NCERT Solutions for Class 10 Social
- NCERT Solutions for Class 10 English
- NCERT Solutions for Class 10 Hindi
- NCERT Solutions for Class 10 Sanskrit
- NCERT Solutions for Class 10 Foundation of IT
- RD Sharma Class 10 Solutions
Formulae Handbook for Class 10 Maths and Science
- Chapter 1 Real Numbers
- Chapter 2 Polynomials
- Chapter 3 Pair of Linear Equations in Two Variables
- Chapter 4 Quadratic Equations
- Chapter 5 Arithmetic Progressions
- Chapter 6 Triangles
- Chapter 7 Coordinate Geometry
- Chapter 8 Introduction to Trigonometry
- Chapter 9 Some Applications of Trigonometry
- Chapter 10 Circles
- Chapter 11 Constructions
- Chapter 12 Areas Related to Circles
- Chapter 13 Surface Areas and Volumes
- Chapter 14 Statistics
- Chapter 15 Probability
We hope the NCERT Solutions for Class 10 Maths Chapter 6 Triangles Ex 6.1, help you. If you have any query regarding NCERT Solutions for Class 10 Maths Chapter 6 Triangles Exercise 6.1, drop a comment below and we will get back to you at the earliest.
Free Resources
Quick Resources
- New QB365-SLMS
- 12th Standard Materials
- 11th Standard Materials
- 10th Standard Materials
- 9th Standard Materials
- 8th Standard Materials
- 7th Standard Materials
- 6th Standard Materials
- 12th Standard CBSE Materials
- 11th Standard CBSE Materials
- 10th Standard CBSE Materials
- 9th Standard CBSE Materials
- 8th Standard CBSE Materials
- 7th Standard CBSE Materials
- 6th Standard CBSE Materials
- Tamilnadu Stateboard
- Scholarship Exams
- Scholarships

CBSE 10th Standard Maths Subject Triangles Case Study Questions 2021
By QB365 on 22 May, 2021
QB365 Provides the updated CASE Study Questions for Class 10 Maths, and also provide the detail solution for each and every case study questions . Case study questions are latest updated question pattern from NCERT, QB365 will helps to get more marks in Exams
QB365 - Question Bank Software
10th Standard CBSE
Final Semester - June 2015
Case Study Questions
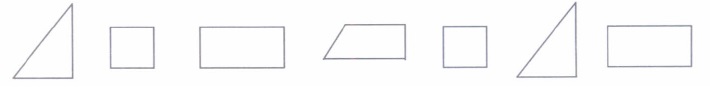

(ii) If m, n and r are the sides of right triangle ABJ, then which of the following can be correct?
| +n = r | +n +r -=0 |
| + n = 2r |
(iii) If \(\Delta\) ABJ ~ \(\Delta\) ADH, then which similarity criterion is used here?
(iv) If \(\angle\) ABJ = 90° and B, J are mid points of sides AD and AH respectively and BJ || DH, then which of the following option is false?
| \((a) \triangle A B J \sim \triangle A D H\) | \((b) 2 B J=D H\) | \((c) A J^{2}=J B^{2}+A B^{2}\) | \((d) \frac{A B}{B D}=\frac{A J}{A H}\) |
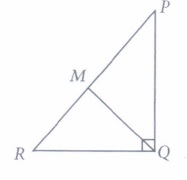
| \((a) \Delta P M Q \sim \Delta P Q R\) |
| \((b) Q R^{2}=P R^{2}-P Q^{2}\) |
| \((c) P R^{2}=P Q+Q R\) |
| \((d) \Delta P M Q \sim \Delta Q M R\) |
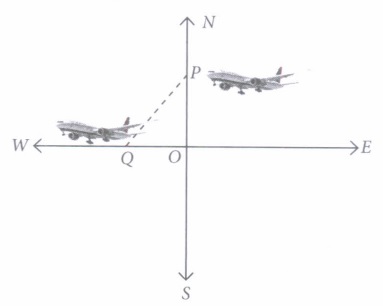
(ii) Distance travelled by aeroplane towards west after \(1 \frac{1}{2}\) hr is
(iii) In the given figure, \(\angle\) POQ is
(iv) Distance between aeroplanes after \(1 \frac{1}{2}\) hr is
| \((a) 450 \sqrt{41} \mathrm{~km}\) | \((b) 350 \sqrt{31} \mathrm{~km}\) | \((c) 125 \sqrt{12} \mathrm{~km}\) | \((d) 472 \sqrt{41} \mathrm{~km}\) |
(v) Area of \(\Delta\) POQ is
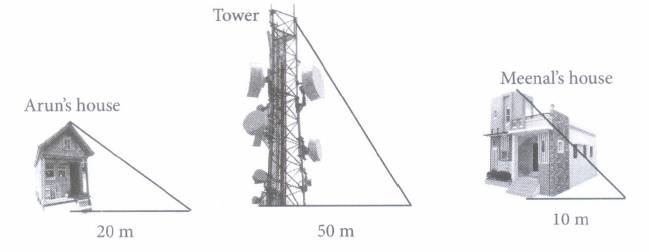
(ii) What will be the length of shadow of tower when Meenal's house casts a shadow of 15 m?
(iii) Height of Aruns house is
(iv) If tower casts a shadow of 40 rn, then find the length of shadow of Arun's house
(v) If tower casts a shadow of 40 m, then what will be the length of shadow of Meenal's house?
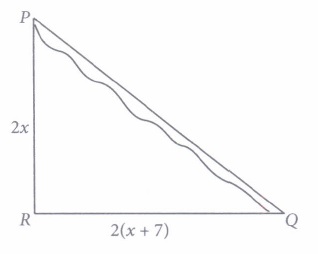
(ii) The value of x is
(iii) The value of PR is
(iv) The value of RQ is
(v) How much distance will be saved in reaching city Q after the construction of highway?
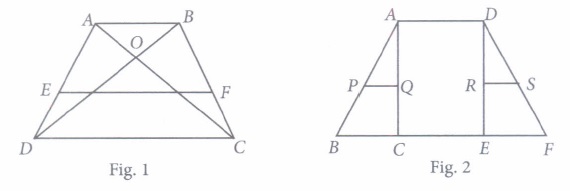
(ii) In if AB || CD, and DO = 3x - 19, OB = x - 5, OC = x - 3 and AO = 3, then the value of x can be
(iii) In if OD = 3x - 1, OB = 5x - 3, OC = 2x + 1 and AO = 6x - 5, then the value of x is
(iv) In \(\Delta\) ABC, if PQ || BC and AP = 2.4 cm, AQ = 2 cm, QC = 3 cm and BC = 6 cm, then AB + PQ is equal to
(v) In \(\Delta\) DEF, if RS || EF, DR = 4x - 3, DS = 8x - 7, ER = 3x - 1 and FS = 5x - 3, then the value of x is
*****************************************
Cbse 10th standard maths subject triangles case study questions 2021 answer keys.
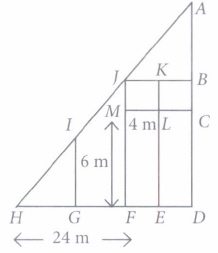
(i) (a): Speed = 1200 km/hr \(\text { Time }=1 \frac{1}{2} \mathrm{hr}=\frac{3}{2} \mathrm{hr}\) \(\therefore\) Required distance = Speed x Time \(=1200 \times \frac{3}{2}=1800 \mathrm{~km}\) (ii) (c): Speed = 1500 km/hr Time = \(\frac{3}{2}\) hr. \(\therefore\) Required distance = Speed x Time \(=1500 \times \frac{3}{2}=2250 \mathrm{~km}\) (iii) (b): Clearly, directions are always perpendicular to each other. \(\therefore \quad \angle P O Q=90^{\circ}\) (iv) (a): Distance between aeroplanes after \(1\frac{1}{2}\) hour \(\begin{array}{l} =\sqrt{(1800)^{2}+(2250)^{2}}=\sqrt{3240000+5062500} \\ =\sqrt{8302500}=450 \sqrt{41} \mathrm{~km} \end{array}\) (v) (d): Area of \(\Delta\) POQ= \(\frac{1}{2}\) x base x height \(=\frac{1}{2} \times 2250 \times 1800=2250 \times 900=2025000 \mathrm{~km}^{2}\)
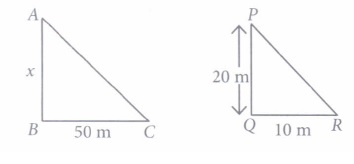
(i) (b) (ii) (c): Using Pythagoras theorem, we have PQ 2 = PR 2 + RQ 2 \(\Rightarrow(26)^{2}=(2 x)^{2}+(2(x+7))^{2} \Rightarrow 676=4 x^{2}+4(x+7)^{2} \) \(\Rightarrow 169=x^{2}+x^{2}+49+14 x \Rightarrow x^{2}+7 x-60=0\) \(\Rightarrow x^{2}+12 x-5 x-60=0 \) \(\Rightarrow x(x+12)-5(x+12)=0 \Rightarrow(x-5)(x+12)=0 \) \(\Rightarrow x=5, x=-12\) \(\therefore \quad x=5\) [Since length can't be negative] (iii) (a) : PR = 2x = 2 x 5 = 10 km (iv) (b): RQ= 2(x + 7) = 2(5 + 7) = 24 km (v) (d): Since, PR + RQ = 10 + 24 = 34 km Saved distance = 34 - 26 = 8 km
(i) (c) (ii) (b): Since \(\Delta\) AOB ~ \(\Delta\) COD [ByAA similarity criterion] \(\therefore \frac{A O}{O C}=\frac{B O}{O D} \Rightarrow \frac{3}{x-3}=\frac{x-5}{3 x-19}\) \(\Rightarrow 3(3 x-19)=(x-5)(x-3) \) \(\Rightarrow 9 x-57=x^{2}-3 x-5 x+15 \Rightarrow x^{2}-17 x+72=0 \) \(\Rightarrow(x-8)(x-9)=0 \Rightarrow x=8 \text { or } 9\) (iii) (c) : Since, \(\Delta\) AOB ~ \(\Delta\) COD [ByAA similarity criterion] \(\therefore \frac{A O}{O C}=\frac{B O}{O D} \Rightarrow \frac{6 x-5}{2 x+1}=\frac{5 x-3}{3 x-1}\) \(\Rightarrow(6 x-5)(3 x=1)=(5 x-3)(2 x+1) \) \(\Rightarrow \quad 18 x^{2}-6 x-15 x+5=10 x^{2}+5 x-6 x-3 \) \(\Rightarrow \quad 8 x^{2}-20 x+8=0 \Rightarrow 2 x^{2}-5 x+2=0\) From options, x = 2 is the only value that satisfies this equation. (iv) (d): Since, \(\Delta\) APQ ~ \(\Delta\) ABC [ByAA similarity criterion] \(\therefore \quad \frac{A P^{\circ}}{A B}=\frac{A Q}{A C}=\frac{P Q}{B C} \Rightarrow \frac{2.4}{A B}=\frac{2}{5}=\frac{P Q}{6} \) \(\therefore \quad A B=\frac{2.4 \times 5}{2}=6 \mathrm{~cm} \text { and } P Q=\frac{2 \times 6}{5}=2.4 \mathrm{~cm} \) \(\therefore \quad A B+P Q=6+2.4=8.4 \mathrm{~cm}\) (v) (a): Since, \(\Delta\) DRS ~ \(\Delta\) DEF [ByAA similarity criterion] \(\therefore \quad \frac{D E}{D R}=\frac{D F}{D S} \Rightarrow \frac{D E}{D R}-1=\frac{D F}{D S}-1 \) \(\Rightarrow \frac{D E-D R}{D R}=\frac{D F-D S}{D S} \Rightarrow \frac{E R}{D R}=\frac{F S}{D S} \) \(\Rightarrow \quad \frac{D R}{E R}=\frac{D S}{F S} \Rightarrow \frac{4 x-3}{3 x-1}=\frac{8 x-7}{5 x-3} \) \(\Rightarrow \quad 20 x^{2}-12 x-15 x+9=24 x^{2}-8 x-21 x+7 \) \(\Rightarrow \quad 4 x^{2}-2 x-2=0 \Rightarrow 2 x^{2}-x-1=0\) Only option (a) i.e., x = 1 satisfies this equation.
Related 10th Standard CBSE Maths Materials
10th standard cbse syllabus & materials, cbse 10th social science the making of a global world chapter case study question with answers, cbse 10th social science nationalism in india chapter case study question with answers, cbse 10th social science the rise of nationalism in europe chapter case study question with answers, cbse 10th science metals and non metals chapter case study question with answers, cbse 10th science acids, bases and salts chapter case study question with answers, cbse 10th science chemical reactions and equations chapter case study question with answers, class 10th science - our environment case study questions and answers 2022 - 2023, class 10th science - magnetic effects of electric current case study questions and answers 2022 - 2023, class 10th science - electricity case study questions and answers 2022 - 2023, class 10th science - human eye and the colourful world case study questions and answers 2022 - 2023, class 10th science - light reflection and refraction case study questions and answers 2022 - 2023, class 10th science - heredity and evolution case study questions and answers 2022 - 2023, class 10th science - how do organisms reproduce case study questions and answers 2022 - 2023, class 10th science - life processes case study questions and answers 2022 - 2023, class 10th science - periodic classification of elements case study questions and answers 2022 - 2023.

Class VI to XII
Tn state board / cbse, 3000+ q&a's per subject, score high marks.

10th Standard CBSE Study Materials

10th Standard CBSE Subjects

NCERT Solutions for Class 10 Maths Chapter 6 Triangles

- Exercise 6.1
- Exercise 6.2
- Exercise 6.3
- Exercise 6.4
- Exercise 6.5
- Exercise 6.6
NCERT Solutions for Class 10 Maths Chapters:
How many exercises in Chapter 6 Triangles
What do you mean by aa or aaa similarity criterion, what is rhs similarity criterion, what is converse of pythagoras theorem, contact form.
NCERT Solutions for Class 10 Maths Chapter 6 Triangles
NCERT Solutions Class 10 Maths Chapter 6 Triangles are provided here to help the students of CBSE class 10. Our expert teachers prepared all these solutions as per the latest CBSE syllabus and guidelines. In this chapter, we have discussed the similarity of triangles, criterion for similarity of triangles, areas of similar triangles and Pythagoras theorem. CBSE Class 10 Maths solutions provide a detailed and step-wise explanation of each answer to the questions given in the exercises of NCERT books.
CBSE Class 10 Maths Chapter 6 Triangles Solutions
Below we have given the answers to all the questions present in Triangles in our NCERT Solutions for Class 10 Maths chapter 6. In this lesson, students are introduced to a lot of important concepts that will be useful for those who wish to pursue mathematics as a subject in their future classes. Based on these solutions, students can prepare for their upcoming Board Exams. These solutions are helpful as the syllabus covered here follows NCERT guidelines.
NCERT Solutions for Class 10 Maths Chapter 6 Exercise 6.1

NCERT Solutions for Class 10 Maths Chapter 6 Exercise 6.2
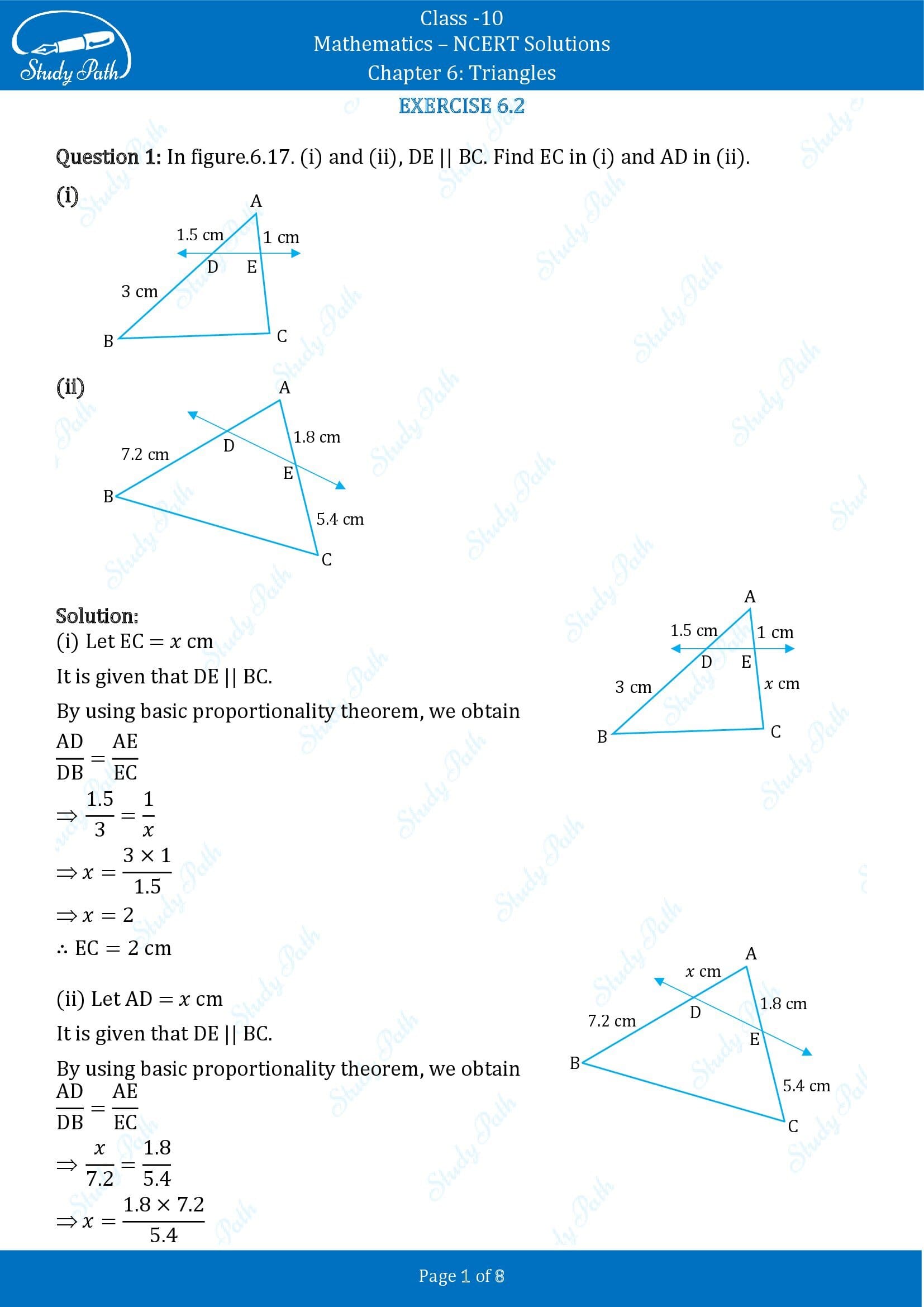
NCERT Solutions for Class 10 Maths Chapter 6 Exercise 6.3
NCERT Solutions for Class 10 Maths Chapter 6 Exercise 6.4
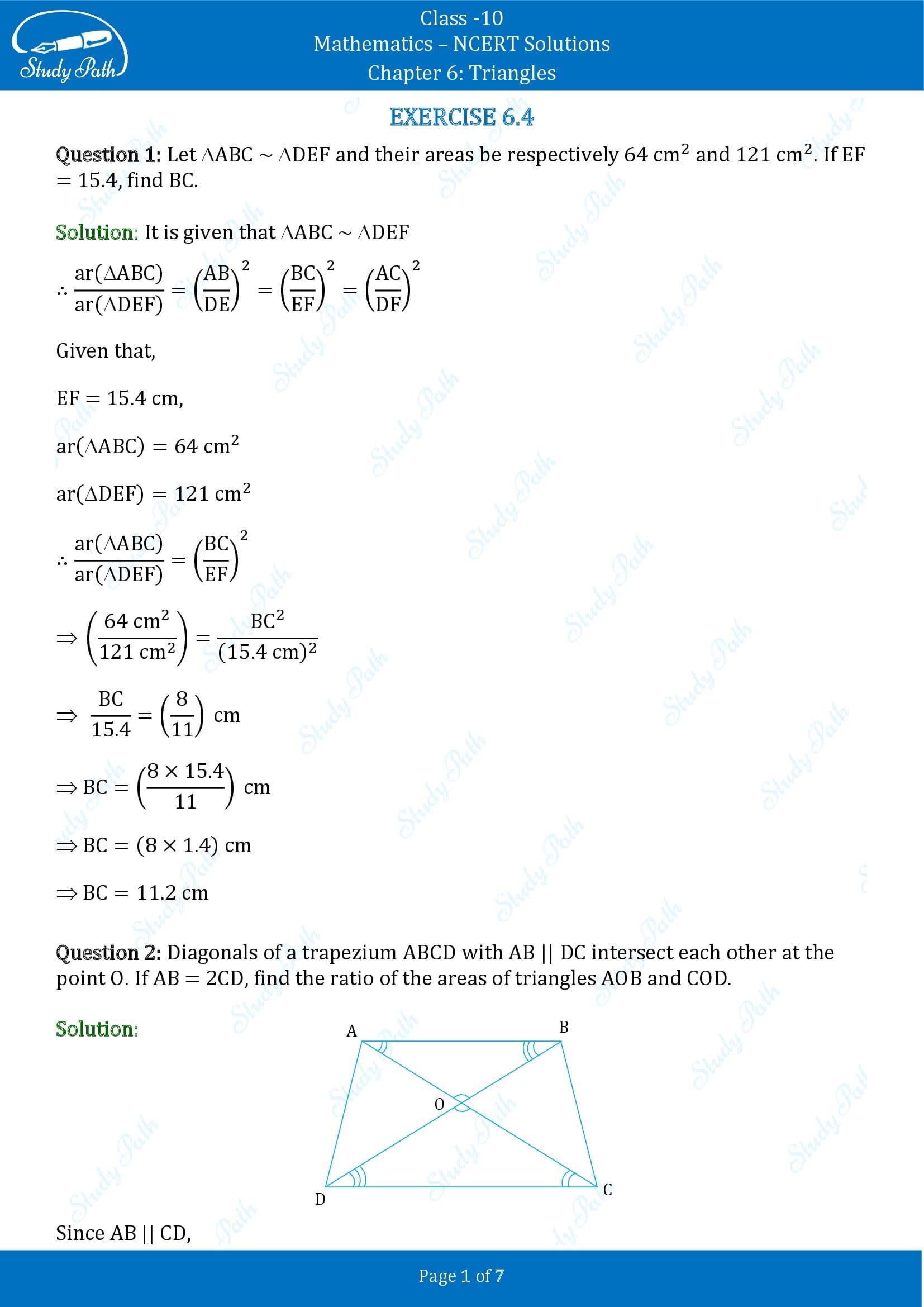

NCERT Solutions for Class 10 Maths Chapter 6 Exercise 6.5

NCERT Solutions for Class 10 Maths Chapter 6 Exercise 6.6
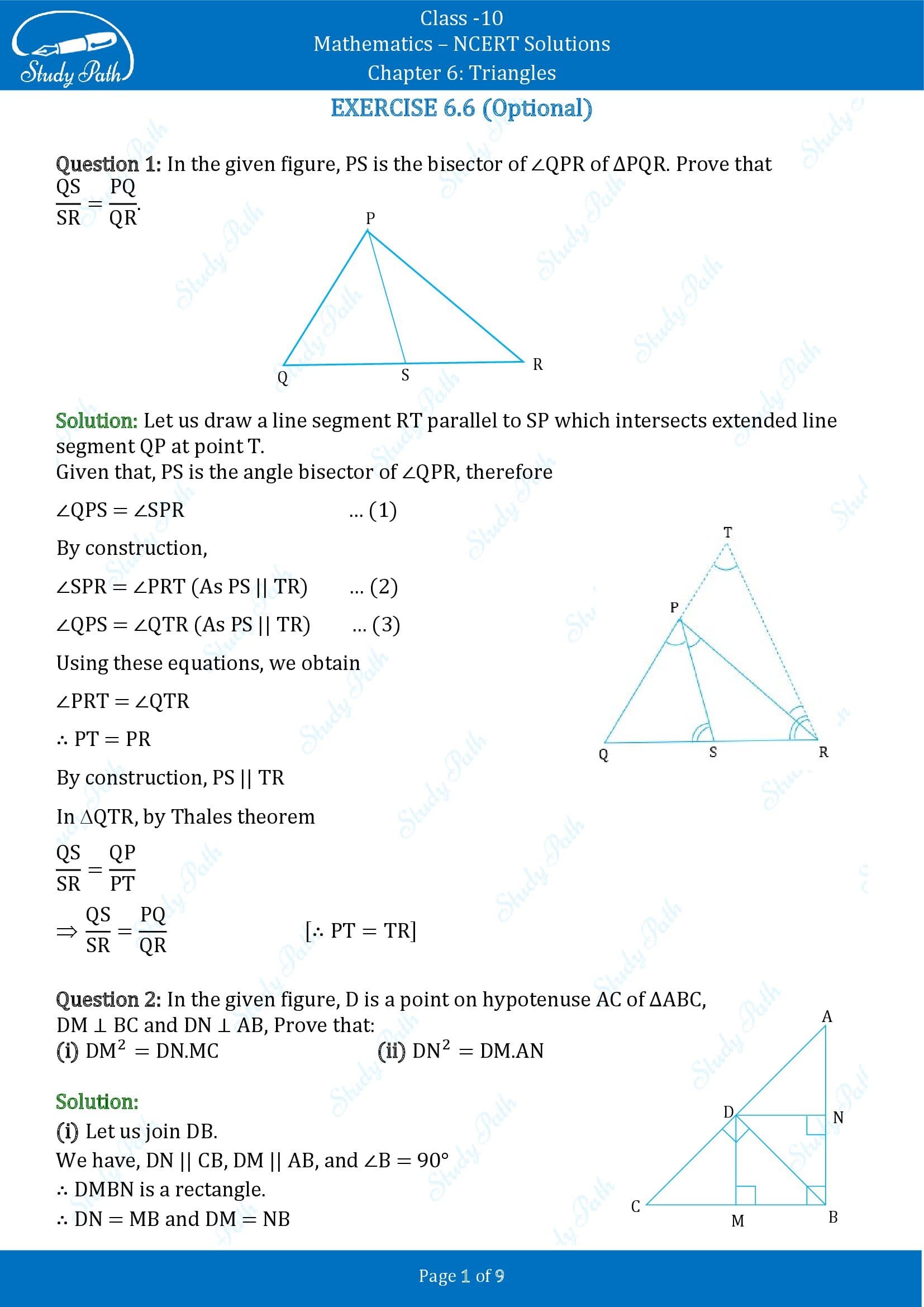

Leave a Reply Cancel reply
Your email address will not be published. Required fields are marked *
Save my name, email, and website in this browser for the next time I comment.
- Class 6 Maths
- Class 6 Science
- Class 6 Social Science
- Class 6 English
- Class 7 Maths
- Class 7 Science
- Class 7 Social Science
- Class 7 English
- Class 8 Maths
- Class 8 Science
- Class 8 Social Science
- Class 8 English
- Class 9 Maths
- Class 9 Science
- Class 9 Social Science
- Class 9 English
- Class 10 Maths
- Class 10 Science
- Class 10 Social Science
- Class 10 English
- Class 11 Maths
- Class 11 Computer Science (Python)
- Class 11 English
- Class 12 Maths
- Class 12 English
- Class 12 Economics
- Class 12 Accountancy
- Class 12 Physics
- Class 12 Chemistry
- Class 12 Biology
- Class 12 Computer Science (Python)
- Class 12 Physical Education
- GST and Accounting Course
- Excel Course
- Tally Course
- Finance and CMA Data Course
- Payroll Course
Interesting
- Learn English
- Learn Excel
- Learn Tally
- Learn GST (Goods and Services Tax)
- Learn Accounting and Finance
- GST Tax Invoice Format
- Accounts Tax Practical
- Tally Ledger List
- GSTR 2A - JSON to Excel
Are you in school ? Do you love Teachoo?
We would love to talk to you! Please fill this form so that we can contact you
CBSE Class 10 Sample Paper for 2021 Boards - Maths Standard
- CBSE Class 10 Sample Paper for 2024 Boards - Maths Standard
- CBSE Class 10 Sample Paper for 2023 Boards - Maths Standard
- Practice Questions CBSE - Maths Class 10 (2023 Boards)
- CBSE Class 10 Sample Paper for 2023 Boards - Maths Basic
- CBSE Class 10 Sample Paper for 2022 Boards - Maths Standard [Term 2]
- CBSE Class 10 Sample Paper for 2022 Boards - Maths Basic [Term 2]
- CBSE Class 10 Sample Paper for 2022 Boards - Maths Standard [MCQ]
- CBSE Class 10 Sample Paper for 2022 Boards - Maths Basic [MCQ]
- CBSE Class 10 Sample Paper for 2021 Boards - Maths Basic
- CBSE Class 10 Sample Paper for 2020 Boards - Maths Standard
- CBSE Class 10 Sample Paper for 2020 Boards - Maths Basic
- CBSE Class 10 Sample Paper for 2019 Boards
- CBSE Class 10 Sample Paper for 2018 Boards
Question 18 (Case Based Question) - CBSE Class 10 Sample Paper for 2021 Boards - Maths Standard - Solutions of Sample Papers for Class 10 Boards
Last updated at April 16, 2024 by Teachoo
Case Study Based- 2 SCALE FACTOR AND SIMILARITY
Scale factor a scale drawing of an object is the same shape as the object but a different size. the scale of a drawing is a comparison of the length used on a drawing to the length it represents. the scale is written as a ratio., similar figures the ratio of two corresponding sides in similar figures is called the scale factor..
Note: Check more Case Based Questions for - Triangles Class 10
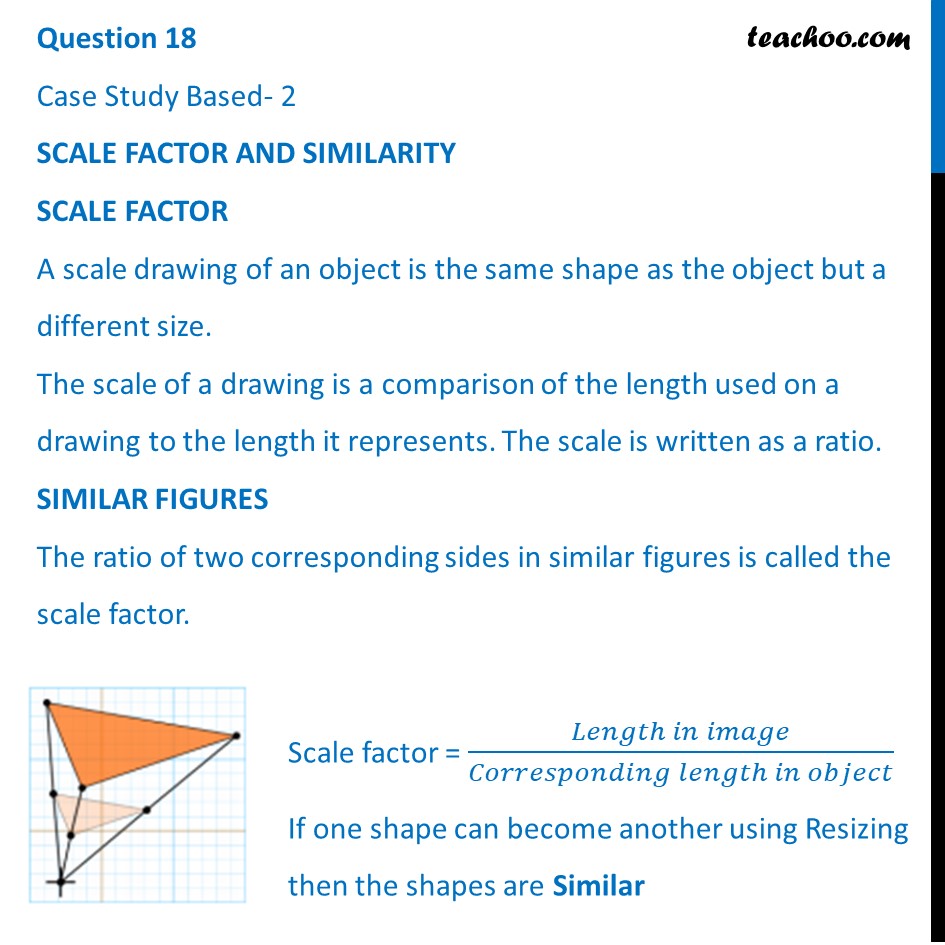
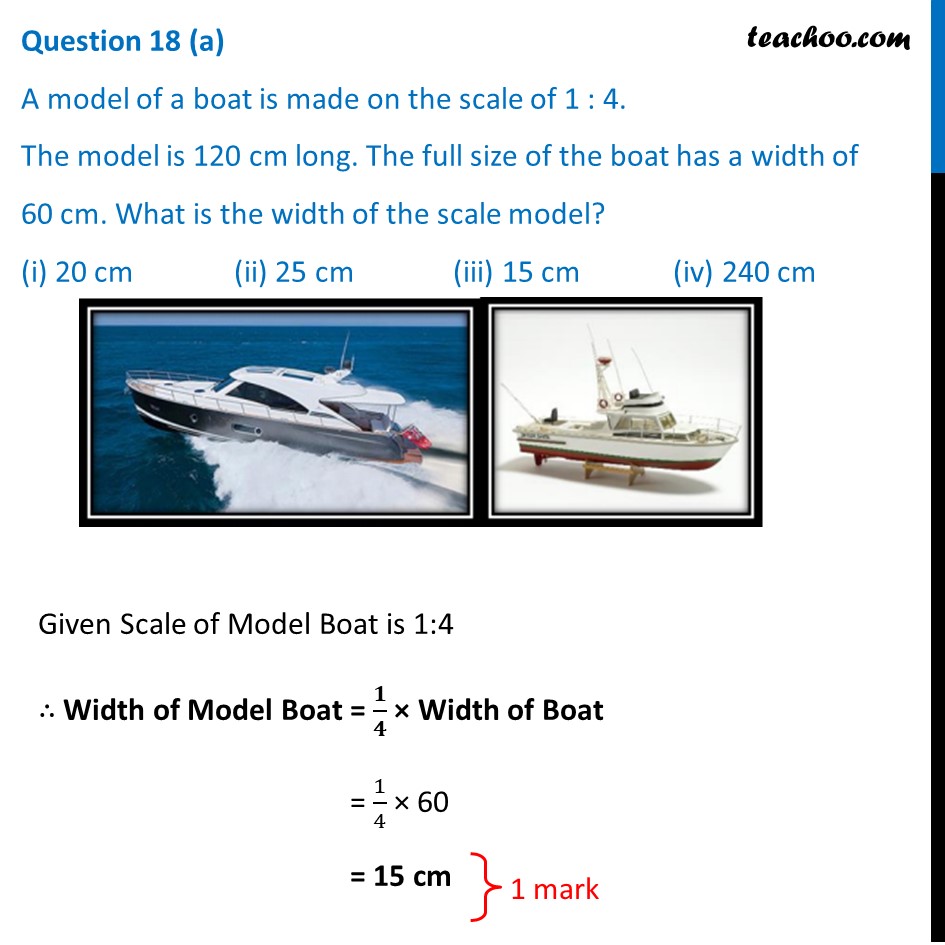






Question 18 Case Study Based- 2 SCALE FACTOR AND SIMILARITY SCALE FACTOR A scale drawing of an object is the same shape as the object but a different size. The scale of a drawing is a comparison of the length used on a drawing to the length it represents. The scale is written as a ratio. SIMILAR FIGURES The ratio of two corresponding sides in similar figures is called the scale factor. Scale factor = (𝐿𝑒𝑛𝑔𝑡ℎ 𝑖𝑛 𝑖𝑚𝑎𝑔𝑒)/(𝐶𝑜𝑟𝑟𝑒𝑠𝑝𝑜𝑛𝑑𝑖𝑛𝑔 𝑙𝑒𝑛𝑔𝑡ℎ 𝑖𝑛 𝑜𝑏𝑗𝑒𝑐𝑡) If one shape can become another using Resizing then the shapes are Similar Rotation or Turn Reflection or Flip Translation or Slide Hence, two shapes are Similar when one can become the other after a resize, flip, slide or turn. (a) A model of a boat is made on the scale of 1 : 4. The model is 120 cm long. The full size of the boat has a width of 60 cm. What is the width of the scale model? (i) 20 cm (ii) 25 cm (iii) 15 cm (iv) 240 cm Given Scale of Model Boat is 1:4 ∴ Width of Model Boat = 𝟏/𝟒 × Width of Boat = 1/4 × 60 = 15 cm (b) What will effect the similarity of any two polygons? (i) They are flipped horizontally (ii) They are dilated by a scale factor (iii) They are translated down (iv) They are not the mirror image of one another We know that triangles are similar if They are flipped They are scaled They are translated Hence, similarity will be effected if they are not the mirror image of one another Hence, similarity will be effected if they are not the mirror image of one another (c) If two similar triangles have a scale factor of a: b. Which statement regarding the two triangles is true? (i) The ratio of their perimeters is 3a : b (ii) Their altitudes have a ratio a : b (iii) Their medians have a ratio 𝑎/2 : b (iv) Their angle bisectors have a ratio a2 : b2If two similar triangles have a scale factor of a: b. Then, Ratio of their altitudes is a:b (d) The shadow of a stick 5 m long is 2 m. At the same time the shadow of a tree 12.5m high is (i) 3 m (ii) 3.5m (iii) 4.5m (iv) 5m Since both these triangles are similar, Their sides are in proportion ∴ (𝑆ℎ𝑎𝑑𝑜𝑤 𝑜𝑓 𝑇𝑟𝑒𝑒)/(𝑆ℎ𝑎𝑑𝑜𝑤 𝑜𝑓 𝑆𝑡𝑖𝑐𝑘)=(𝐿𝑒𝑛𝑔𝑡ℎ 𝑜𝑓 𝑡𝑟𝑒𝑒)/(𝐿𝑒𝑛𝑔𝑡ℎ 𝑜𝑓 𝑠𝑡𝑖𝑐𝑘) (𝑆ℎ𝑎𝑑𝑜𝑤 𝑜𝑓 𝑇𝑟𝑒𝑒)/2=12.5/5 Shadow of Tree = 2 × 12.5/5=𝟓 𝒎 (e) Below you see a student's mathematical model of a farmhouse roof with measurements. The attic floor, ABCD in the model, is a square. The beams that support the roof are the edges of a rectangular prism, EFGHKLMN. E is the middle of AT, F is the middle of BT, G is the middle of CT, and H is the middle of DT. All the edges of the pyramid in the model have length of 12 m. What is the length of EF, where EF is one of the horizontal edges of the block? (i) 24m (ii) 3m (iii) 6m (iv) 10mHere, Δ TEF is similar to Δ TAB Since both these triangles are similar, Their sides are in proportion ∴ 𝑇𝐸/𝑇𝐴=𝐸𝐹/𝐴𝐵 6/12=𝐸𝐹/12 EF = 6 m

Davneet Singh
Davneet Singh has done his B.Tech from Indian Institute of Technology, Kanpur. He has been teaching from the past 14 years. He provides courses for Maths, Science, Social Science, Physics, Chemistry, Computer Science at Teachoo.
Hi, it looks like you're using AdBlock :(
Please login to view more pages. it's free :), solve all your doubts with teachoo black.
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Open access
- Published: 15 July 2024
A One Health framework for exploring zoonotic interactions demonstrated through a case study
- Amélie Desvars-Larrive ORCID: orcid.org/0000-0001-7671-696X 1 , 2 ,
- Anna Elisabeth Vogl ORCID: orcid.org/0009-0007-9813-3854 1 ,
- Gavrila Amadea Puspitarani ORCID: orcid.org/0000-0002-8826-5606 1 , 2 ,
- Liuhuaying Yang 2 ,
- Anja Joachim ORCID: orcid.org/0000-0003-3082-6885 3 &
- Annemarie Käsbohrer ORCID: orcid.org/0000-0003-2966-2713 1
Nature Communications volume 15 , Article number: 5650 ( 2024 ) Cite this article
1565 Accesses
138 Altmetric
Metrics details
- Bacterial infection
- Ecological networks
- Epidemiology
- Parasitic infection
- Viral infection
The eco-epidemiology of zoonoses is often oversimplified to host-pathogen interactions while findings derived from global datasets are rarely directly transferable to smaller-scale contexts. Through a systematic literature search, we compiled a dataset of naturally occurring zoonotic interactions in Austria, spanning 1975–2022. We introduce the concept of zoonotic web to describe the complex relationships between zoonotic agents, their hosts, vectors, food, and environmental sources. The zoonotic web was explored through network analysis. After controlling for research effort, we demonstrate that, within the projected unipartite source-source network of zoonotic agent sharing, the most influential zoonotic sources are human, cattle, chicken, and some meat products. Analysis of the One Health 3-cliques (triangular sets of nodes representing human, animal, and environment) confirms the increased probability of zoonotic spillover at human-cattle and human-food interfaces. We characterise six communities of zoonotic agent sharing, which assembly patterns are likely driven by highly connected infectious agents in the zoonotic web, proximity to human, and anthropogenic activities. Additionally, we report a frequency of emerging zoonotic diseases in Austria of one every six years. Here, we present a flexible network-based approach that offers insights into zoonotic transmission chains, facilitating the development of locally-relevant One Health strategies against zoonoses.
Similar content being viewed by others
The science of the host–virus network
Micro-level social structures and the success of COVID-19 national policies
A geopositioned and evidence-graded pan-species compendium of Mayaro virus occurrence
Introduction.
Zoonoses are caused by pathogens naturally transmissible between humans and wild or domestic animals. Places where humans and animals or animal products interact create interfaces that facilitate zoonotic agent transmission. Notably, approximately 99% of endemic zoonotic infections in humans originate from domesticated animals, within anthropogenic environments, either directly or indirectly through contaminated food or vectors 1 . Morand, et al. 2 provided statistical evidence supporting the positive relationship between the duration of domestication and the diversity of zoonotic agents that humans share with each domestic species, which was initially hypothesised by McNeill 3 . In addition, over 60% of human emerging infectious diseases (EIDs) are zoonotic 4 . Although direct zoonotic spillover from wildlife is rare and wildlife-to-human transmission typically occurs through indirect transmission 1 , more than 70% of these zoonotic emergences are caused by pathogens with a wildlife origin 4 . However, the full host breadth of endemic and emerging zoonotic agents, as well as their animal and environmental reservoirs are rarely identified nor mapped.
In most zoonotic disease systems, interactions occur among multiple animal host species, environmental sources (including invertebrate vectors), and involve multiple infectious agents 5 . Therefore, exploring disease dynamics in these multi-source, multi-agent systems necessitates considering the complex ecology of the interactions, e.g., the host-pathogen community assemblages, the existence of environmental reservoirs, and the involvement of vectors 5 , 6 , 7 . Unfortunately, this complexity is often ignored due to the lack of comprehensive datasets, making it challenging to embrace a transdisciplinary perspective. Furthermore, network approaches to infectious diseases and spillover risk have largely focused on the analysis of the host-pathogen relationships 2 , 8 , 9 , 10 , 11 , neglecting other sources of zoonotic infection, such as contaminated environment or food. A comprehensive understanding of circulating zoonotic agents, their hosts, vectors, food and environmental sources, and the key interfaces where spillover events may occur is essential for developing effective integrated One Health monitoring, prevention, and control of zoonoses 12 .
Zoonotic and emerging diseases pose a significant threat to both human and animal health 13 , 14 , they cause substantial economic losses 15 , and may have far-reaching consequences on multiple aspects of the society 16 . The enhancement of monitoring efforts and data collection in both domestic and wildlife hosts is essential for effectively predicting the establishment of reservoirs, understanding the facilitators of zoonotic spillover, and preventing such spillover at source 17 . However, the ecology and diversity of circulating zoonotic agents are tied to multiple factors, including the local availability of potential animal hosts and vectors, their spatial distribution, density, population dynamics, and community composition 18 , 19 . In addition, the spillover force of infection depends on cultural and socio-economic determinants, including human agricultural practices, feeding and hunting habits, and behaviour 20 , 21 . This underscores the pressing need for the development of analytical tools to optimise surveillance strategies that are tailored to the regional or national context. While global datasets may be available 8 , 22 , 23 , 24 , data granularity and completeness are generally suboptimal for smaller-scale investigations. Furthermore, there is a scarcity of national studies focusing on zoonotic interfaces that encompass animals, vectors, environmental, and food matrices. Bridging this gap is crucial for developing effective, locally relevant strategies 25 to monitor and mitigate potential changes in spillover risk that could impact human and animal health.
Austria has a growing population of nine million people. Its fauna encompasses approximately 45,870 species, of which 626 are vertebrates, including 110 mammalian and 418 avian species 26 . Moreover, of 3.9 million Austrian households, 35% own pets. The country also counts ~ 53,300 cattle, one million pigs, and five million poultry, while 133,000 hunting permits are issued annually 27 . These numbers underline the importance of the human-animal interfaces at the national scale. Given the potential for zoonotic disease transmission at these interfaces and the ensuing risk to human health, Austria adheres to a combination of European and national regulations, guaranteeing a framework for coordinated epidemiological surveillance and responses. However, concentrating mostly on notifiable diseases, monitored and reported only for specific species, official figures tend to overlook non-regulated zoonotic agents circulating in the territory that could pose a risk to public health.
In this study, we extracted data from scientific papers and national laboratory reports spanning 47 years of publications, to generate a real-world network describing the web of zoonotic interactions in Austria and characterise the various interfaces through which zoonotic spillover may occur. We introduce the concept of “zoonotic web” (akin “food web”) 28 as a network representation of zoonotic actors at human-animal-environment interfaces (i.e., [host-vector-environment-food]-zoonotic agent network), intended for use in One Health approaches. We treated it as a bipartite network and transformed it into a one-mode projection representing the network of zoonotic agent sharing among zoonotic sources, weighting relationships (edges) between zoonotic sources (nodes) by the number of zoonotic agents they shared. We explored this network using different network centrality metrics and a community-based approach. In addition, we examined zoonotic disease emergence patterns in Austria and pinpointed research trends and gaps on zoonotic agents at the national level.
Dataset of zoonotic interactions in Austria, 1975–2022
The search identified 2186 publications. After 542 duplicates were removed, 1644 publications were screened with 1269 excluded at the title/abstract screening stage as they were not eligible (see Supplementary Fig. 1 for a breakdown based on exclusion criteria). This left 375 publications, of which 16 could not be retrieved, so 359 full-text articles were assessed for eligibility, with 229 meeting the criteria for final inclusion. In addition, 17 publications were found in excluded review articles, leading to a total of 246 publications that were ultimately included in this study (168 scientific articles, 13 reports, and 65 theses).
The final dataset is a *.csv. file with 2128 rows and 48 data fields. Each row represents one investigated zoonotic agent along with the results of the investigation in the animal host(s), vector(s), and environmental or food matrix(-ices). All included publications were published between January 1975 and August 2022. We evidenced a 17.8-fold increase in the number of publications on zoonoses in Austria between the first (1975–1997) and the second half (1998–2022) of the study period (Supplementary Fig. 2 ). In addition, there was variation in study distribution among federal states (Supplementary Fig. 3 ). To contextualise this result, it was compared with global data: a PubMed search using the terms (zoono* OR “zoono* disease*“) from 1975 until 23 August 2022 (without restricting the search to Austria) generated a total of 64,282 results and revealed an increase of the same order (~ 18-fold). However, a PubMed search using the term “health” in the same period yielded 5,791,763 results, indicating a mere 6.8-fold increase in health publications globally. This result suggests a disproportionate rise in zoonotic disease research, both at the national and international levels, compared to general health studies.
Research trends
Between 1975 and 2022, 227 unique zoonotic agents were investigated in Austria (not all of them could be resolved at species level). Ten genera collectively accounted for 41% of the selected literature: Salmonella, Escherichia, Listeria, Echinococcus, Orthoflavivirus, Brucella, Toxoplasma, Campylobacter, Trichinella , and Leptospira (Supplementary Table 1 ). Most zoonotic agents were studied in wildlife hosts, which accounted for 76.9% of the 221 animal species investigated. Furthermore, during the study period, the majority of investigations into food products concentrated on animal-origin products whereas plant-based foods (including fruits, vegetables, spices/herbs, and grains) accounted for 5.6% of the examined foodstuffs. Finally, across the selected publications, seven environmental matrices (including food and processing plants, public lavatory, sandbox, slaughter knife, soil, and water) and 21 invertebrate taxa (mosquitoes: 47.8%; ticks; 39.1%; sand flies, gastropods, and fleas: 4.3% each) were investigated.
In Austria, there has been a noticeable upward trend in scientific interest regarding zoonotic bacteria, viruses, and eukaryotes (Fig. 1a ), with bacteria garnering the most attention. We observed an upward trend across all compartments, as recognised by the traditional One Health triad, i.e., animal, human, and environment, followed by a subsequent decrease in the number of studies investigating animals (from 2015) and humans (from 2010). The environmental aspect (including environmental media, plant-based food, and vectors) was not considered in studies on zoonotic diseases in Austria until 1997 but subsequently demonstrated the most gradual increase in scientific interest (Fig. 1b ), primarily driven by a rise in investigations on vectors (Supplementary Fig. 4 ).
a Trends in research interest measured by the number of investigations involving different superkingdoms of zoonotic agents. b Trends in research interest measured by the number of investigations involving each compartment, as recognised in the traditional One Health view. Dots represent the number of investigations per year; solid lines show a fitted trend (loess regression); shaded areas represent the corresponding 95% confidence interval. Only publications that investigated naturally occurring zoonotic infections were considered. Plant-based foodstuffs, invertebrate vectors, and any environmental matrices (including from food processing plants) were grouped under the compartment “environment” while food products of animal origin were considered within the “animal” compartment. Note that a single publication may present more than one investigation, i.e., investigating multiple zoonotic agents belonging to different superkingdoms and/or multiple compartments.
Zoonotic web actors and interfaces
Overall, between 1975 and 2022, the literature reported 197 zoonotic agents in Austria that were directly or indirectly evidenced in natural infections, including an unusual case of a dog hair described as a zoonosis (this “agent” was not considered in the network analysis) (Supplementary Fig. 5 ). Among them, 187 (94.9%) were directly or indirectly detected in 155 distinct vertebrate hosts, including human, 111 wildlife, eight livestock, and 36 companion animal (including exotic pets) species (Supplementary Table 2 ). The highest zoonotic agent richness was observed in Primates (88, with 87 zoonotic agents reported in humans), Carnivora (59 zoonotic agents), Artiodactyla (59), Galliformes (24), and Rodentia (23) (Fig. 2 ). In 78.6% (777 out of 989) of the positive results in hosts, direct detection was achieved and represented the preferred method for bacteria and eukaryotes across all investigated host taxonomic classes. Conversely, viral circulation was primarily evidenced by indirect methods detecting antibodies (Table 1 ).

a Bubble plot illustrating the distribution of the zoonotic agent genera across Austrian hosts grouped by order, 1975–2022. Only publications that investigated naturally occurring zoonotic infections were considered. Bubble size corresponds to the number of zoonotic agents detected within a specific genus during this timeframe. Colours correspond to the zoonotic agent superkingdom. b Zoonotic agent richness per host taxonomic class, with data disaggregated for human and non-human mammals. c Zoonotic agent richness per host taxonomic order. When the host scientific or common name was not specific enough within the publication, the taxonomic order could not be retrieved, and the host name, as mentioned in the publication, was used (e.g., lizard, new-world camelid).
At the environment-zoonotic agent interface, 24 (12.2%) zoonotic agents were detected in 12 different invertebrate (vector) species. Surprisingly, despite the detection of the Usutu virus (USUV) in various bird species, horses, and humans across the reviewed studies, it was not reported in arthropod vectors, a necessary component of its biological cycle (Supplementary Table 3 ). In addition, 11 (5.6%) zoonotic agents, including bacteria ( Listeria monocytogenes , Salmonella sp., Escherichia coli , and Mycobacterium sp.) and eukaryotes ( Cryptosporidium, Giardia , and Toxocara ) were reported in six types of environmental media, including surfaces and tools in food processing environments as well as “natural” matrices (e.g., water, sandboxes) (Supplementary Table 4 ). Finally, at the food system-zoonotic agent interface, 15 (7.6%) zoonotic agents were detected in 31 categories of food. Meat and meat products yielded the majority of positive results (55.8%) while plant-based foods comprised only 2.5% of zoonotic agent-positive food products. Zoonotic agents identified in food were mainly of the genera Listeria (36.6% of positive foodstuffs), Escherichia (22.8%), and Salmonella (22.5%). Out of the 21 identified zoonotic agents in foodstuffs, all were bacteria except for three parasites ( Anisakis, Echinococcus , and Trichinella spiralis ) (Supplementary Fig. 6 ).
Zoonotic web structure
Figure 3 depicts the zoonotic web and interfaces (see Supplementary Fig. 7 for conventional bipartite network visualisation). The network contained 396 nodes, i.e., actors (zoonotic sources and agents), with 658 edges (representing infections), and an average number of 1.66 interactions per actor. The giant connected component of the zoonotic web included 387 actors (97.7% of the nodes) with 652 edges (99.1% of the edges). In addition, the zoonotic web comprised three small components: the first illustrated relationships between Encephalitozoon cuniculi and its hosts Arvicola amphibius (Eurasian water vole) and Oryctolagus cuniculus (rabbit); the second showed Mycobacterium chelonae , Mycobacterium marinum , and Mycolicibacterium fortuitum with their common host Salmo trutta fario (river trout); finally, the third depicted the infection of mosquitoes of the genus Uranotaenia with Alphamesonivirus 1.
This representation uses the D3 forceLink layout, providing a detailed visualisation and offering epidemiological insights into naturally occurring zoonotic interactions in Austria. The zoonotic web is a bipartite network, where each node (circle) represents an actor in the zoonotic web, with one set of nodes representing zoonotic agents (black nodes) and the second set representing zoonotic sources that belong to different categories: vertebrate hosts (red nodes), vectors (yellow nodes), foodstuffs (blue nodes), and environmental matrices (green nodes). A link between a zoonotic agent i and a vertebrate host j indicates that agent i was directly or indirectly detected in host j ; a link between a zoonotic agent i and a vector j signifies that agent i was identified in vector j , implying that vector j may transmit agent i to a vertebrate host through a bite or mechanically; a link between a zoonotic agent i and an environmental matrix j indicates the presence of agent i in environment j , potentially leading to infection of a vertebrate host upon contact ; and a link between a zoonotic agent i and a food matrix j indicates that agent i was detected in food j , which may result in the infection of a vertebrate host through ingestion. Node size represents the actor’s degree centrality. The node degree centrality for each zoonotic source corresponds to the zoonotic agent richness, i.e., the number of taxa directly or indirectly evidenced from the zoonotic source. The node degree centrality for each zoonotic agent corresponds to the zoonotic source range, i.e., the number of sources from which the agent has been directly or indirectly evidenced, reflecting its “host” or “zoonotic source” plasticity. The bottom-right graph illustrates the degree distribution for the “zoonotic agents” and “zoonotic sources” partitions, the latter being disaggregated based on source categories. Interactive version at: https://vis.csh.ac.at/zoonotic-web/dashboard.html .
The analysis of the zoonotic web showed a right-skewed distribution of the node degree centrality (the number of links a node has), revealing few nodes with a high number of connections whereas most of the nodes had one. Among the hosts, the nodes Homo sapiens (human, degree centrality, k = 87), Bos taurus (cattle, k = 38), Canis lupus familiaris (dog, k = 29), Felis catus (domestic cat , k = 21), Vulpes vulpes (red fox, k = 19), Sus scrofa (pig, k = 17), Gallus gallus (chicken, k = 15), Ovies aries (sheep, k = 13), Sus scrofa (w) (wild boar, k = 11), and Nyctereutes procyonoides (raccoon dog, k = 10) exhibited high zoonotic agent richness. Among the vectors, the node Ixodes exhibited the highest degree centrality ( k = 16), with multiple connections to Rickettsia, Borrelia , and Babesia species. In contrast, the node Culex showed a low degree centrality ( k = 2), with links to West Nile virus (WNV) and Orthobunyavirus Tahyna. Among nodes representing food sources, the highest degree centrality was observed for the nodes cattle meat and meat product , animal (unspecified) meat and meat product (each k = 8), and animal (unspecified) dairy ( k = 6). The degree centrality of nodes representing environmental matrices showed relatively low values, ranging between 1 and 4. Among the zoonotic agents, the nodes USUV ( k = 38) , Salmonella enterica ( k = 33), WNV ( k = 30), Salmonella ( k = 24), Escherichia coli ( k = 19), Listeria ( k = 17), Listeria monocytogenes ( k = 17), verotoxigenic Escherichia coli (VTEC) ( k = 16), Campylobacter jejuni ( k = 15), Toxoplasma gondii ( k = 15), Influenza A virus ( k = 12), Campylobacter coli ( k = 11), enterohaemorrhagic E. coli (EHEC) ( k = 11), Leptospira , Staphylococcus aureus , and Campylobacter (each k = 10) revealed a greater zoonotic source plasticity (range) in Austria. Furthermore, with an average degree of 3.90, viruses had a greater zoonotic source plasticity than bacterial (3.77) or eukaryotic (2.28) zoonotic agents.
Network of zoonotic agent sharing
We generated a unipartite scientific research effort-adjusted network of zoonotic sources (i.e., accounting for research biases), based on zoonotic agent sharing. This network depicts patterns of zoonotic transmission potential between sources, with edges representing the likelihood that a given zoonotic source will transmit one or more zoonotic agents to another source relative to other sources in the network 29 . Thus, for one zoonotic agent, connected sources belong to the same potential transmission chain 29 , 30 (Fig. 4a ). In this network, node rankings using the four centrality metrics (degree; strength, i.e., the sum of the weights of edges to/from a node; betweenness, i.e., the number of shortest paths that go through a node; and closeness, i.e., the average distance to all other nodes) 31 showed positive correlation (0.26 < Kendall’s Tau < 0.77, p < 0.001 in all cases, Supplementary Table 5 ). Degree and strength centrality reflect co-occurrence patterns of zoonotic agents among sources 9 . In contrast, betweenness and closeness centrality provide insights into indirect interactions through other sources 32 . The nodes Homo sapiens (human), Gallus gallus (chicken), Bos taurus (cattle), and animal (unspecified) meat and meat product were the most influential nodes in the network, appearing in the top 10 actors by the four centrality metrics. In addition, the nodes Ovies aries (sheep) and cattle meat and meat product could also be considered influential, ranking in the top 10 actors by three (out of four) centrality metrics (Table 2 ). Notably, the nodes Equus caballus (horse) and various nodes representing bird species exhibited high degree and strength centrality, attributable to their shared interactions with the two Orthoflaviviruses, WNV and USUV. Interestingly, the nodes Sus scrofa (wild boar), Testudines (turtles), Canis lupus familiaris (dog), Felis catus (domestic cat), Apodemus flavicollis (yellow-necked field mouse), Nyctereutes procyonoides (raccoon dog), and the tick Ixodes ranked high by betweenness centrality, suggesting that they may act as bridges between host communities 33 , 34 . Besides the most influential nodes, two hosts, Sus scrofa (pig) and Canis lupus familiaris (dog), as well as two food matrices, animal (unspecified) ready to eat product and pig meat and meat product, ranked in the top 10 actors by closeness centrality. Closeness centrality identifies nodes that are “close” to many other nodes 31 ; therefore, zoonotic sources which share numerous zoonotic agents with numerous sources would have high closeness centrality 33 . Summary statistics for the four-node centrality metrics per category of zoonotic sources are shown in Table 3 . Many nodes in the network showed a betweenness equal to zero. Except for betweenness centrality, there were significant differences in the average values of the centrality metrics between the four zoonotic source categories (Supplementary Table 6 ).

The network is unipartite, and each node (circle) represents a zoonotic source that belongs to a source category: vertebrate host, vector, food, or environmental matrix. Node size represents the zoonotic source’s degree. The weight (width) of an edge between two zoonotic sources represents the number of zoonotic agents shared, adjusted for the scientific research effort. a Transmission-potential network among zoonotic sources. Node colours depict zoonotic source categories. Zoonotic hosts are additionally colour-coded based on taxonomic classes to offer further biological insights. b Communities of zoonotic sources based on zoonotic agent sharing as determined using the Leiden algorithm. Node colours represent the communities.
Zoonotic agent sharing communities
We identified six communities (clusters of zoonotic sources sharing similar agents) in the zoonotic agent sharing network (Fig. 4b ). Community 1: primarily comprised of central hosts having higher values of centrality in the unipartite zoonotic agent sharing network and generally living in proximity to humans or having frequent interactions with humans, including livestock, companion animals (dog, cat), synanthropic species (Norway rat, domestic mouse), game species (red fox, cervids), but also captive primates. Notably, Aedes mosquitoes and ticks ( Hyalomma, Ixodes ) were part of this community. Community 1 was characterised by a high diversity of zoonotic agents, with 175 taxa shared among 51 zoonotic sources that composed the community. Community 2: encompassed diverse reptiles (snakes, lizards, and turtles) and amphibians, including non-traditional pet (NTP) species, along with the wild boar; the main zoonotic agent shared within this community was S. enterica . Community 3: consisted of various avian taxa, including birds of prey, ducks, waterfowl, gamebirds, chickens, and pigeons. Note that hosts in this community were broadly designated, lacking specific scientific nomenclature. The primary shared zoonotic agents in community 3 were E. coli and Influenza A virus. Community 4: included various food products and environmental matrices related to food production, but also public lavatory and Meleagris gallopavo (turkey). The main zoonotic agents shared within community 4 were foodborne, principally Salmonella, Listeria monocytogenes , and VTEC. Community 5: mostly clustered WNV hosts and, to a lesser extent, USUV hosts, including various bird species, the vector Culex , and horses. Community 6: represented USUV hosts and exclusively included bird species (Fig. 5 , Supplementary Table 7 and Supplementary Fig. 8 ).

Communities (represented by squares) were determined using the Leiden algorithm run on the research effort-adjusted zoonotic agent sharing network. Circles represent zoonotic agents. The size of each circle represents the degree centrality of the node in the bipartite zoonotic web (i.e., the total number of sources from which it has been directly or indirectly evidenced, reflecting its “host” or “zoonotic source” plasticity). The circulation of a zoonotic agent within a community is represented by a link between the community (square) and the zoonotic agent (circle). Link width represents the number of zoonotic sources that share the zoonotic agent within the community it is linked to. The colour scale shows the number of zoonotic sources (the colour scale is correlated to both node size and link width). Interactive version at: https://vis.csh.ac.at/zoonotic-web/dashboard.html .
Zoonotic agent sharing at human-animal-environment interfaces
A total of 24,475 3-cliques were identified, of which 153 were One Health 3-cliques. The distribution of the research effort-adjusted number of zoonotic agents shared at human-animal-environment interfaces (represented by the sum of the edge weights within One Health cliques) displayed a right-skewed pattern (Fig. 6a ), with a median of 20.8. This suggested that, at most human-animal-environment interfaces, the likelihood of a specific zoonotic source transmitting one or more zoonotic agents to another source is relatively low. We identified six One Health cliques that ranked the highest based on the number of zoonotic agents shared (Fig. 6b ). In five of them, cattle ( B. taurus ) was involved, while in two of them, foodstuffs from animal or plant origin were implicated. Environmental samples from food processing facilities were present in three cliques.

a Histogram and density plot of the total edge weight in One Health 3-cliques within the network of zoonotic agent sharing. The edge weight between two nodes (representing zoonotic sources belonging to different One Health compartments) is the number of zoonotic agents shared between these two nodes, adjusted for the scientific research effort. The total edge weight of a clique corresponds to the sum of the edge weights between all pairs of nodes within the clique. Plant-based foodstuffs, invertebrate vectors, and any environmental matrices (including from food processing plants) were included into the compartment “environment” while food products of animal origin were considered within the “animal” compartment. The dashed line separates the top six cliques (right side) from the others. b Top six One Health 3-cliques based on the total edge weight. Nodes are colour-coded based on the “traditional” One Health compartment, with yellow representing the human compartment, blue representing the animal compartment, and green representing the environment. Edge weight is visualised as the edge width and corresponding edge label.
Imported and emerging zoonotic agents
Between 1975 and 2022, Austria reported 48 importation events of zoonotic agents, of which, 11 were bacteria, seven were helminths, one was an arthropod, and two were viruses. These imported zoonotic agents were documented as potentially originating from multiple countries (Supplementary Table 8 ). In addition, we report the emergence of eight zoonotic diseases in Austria between 1975 and 2022, corresponding to a frequency of approximately one emerging zoonotic disease every six years. Notably, all of them were found to emerge in the past 20 years. The etiologic agents and their respective hosts, along with the year of discovery were: USUV (birds, 2001), Rickettsia helvetica ( Ixodes ricinus , 2005), Anisakis (human, 2009), Brucella canis (dog, 2010), Rickettsia conorii subsp. raoultii (dog, 2015), WNV (horse, 2016), Thelazia capillipaeda (domestic cat, 2018), and Baylisascaris procyonis (racoon, 2019). We documented three types of emergences: the first discovery outside the historical geographic range, the first discovery in Austria, and the first autochthonous case (Supplementary Fig. 9 ). Using the zoonotic web, we additionally estimated the current source range of the eight emerging zoonotic agents, revealing associations with 59 vertebrate hosts, including human, and four genera of arthropod vectors (Supplementary Fig. 10 ).
Cross-species transmission and emergence of zoonotic-origin diseases occur at complex animal-human-environment interfaces, within dynamic social-ecological systems influenced by human behaviour, demographic shifts, and global changes. These interfaces represent significant One Health challenges. Here, we present the first attempt to analyse nearly 50 years of data on naturally occurring zoonotic infections (or contaminations) in Austria, leveraging an original One Health approach based on network theory. With approximately 80% of detections in animals and all those in vectors, food, and environmental matrices supported by direct evidence of zoonotic agents, we are confident in the robustness of our results. This work demonstrates that most zoonotic agents are capable of infecting both human and diverse animal species across various taxa, while evolving within multi-source, multi-agent ecological communities, consistent with the established principles in parasite community ecology 35 . We argue that the analysis of the zoonotic web holds greater value when studying potential zoonotic transmission chains compared to the commonly employed host-pathogen network approach, as it offers a broader epidemiological perspective and more analytical flexibility. Notably, we studied the centrality of zoonotic sources, including hosts, vectors, foodstuffs, and environmental matrices, within the network of zoonotic agent sharing, and evidenced that certain sources play a disproportionate role in the sharing of zoonotic agents. Specifically, we underscored the crucial role of arthropod vectors and foodstuffs (typically omitted in host-pathogen networks) in the risk of zoonotic disease emergence and transmission through the zoonotic web, pinpointing potential targets for One Health surveillance programmes.
Ten genera of zoonotic agents constituted 41% of the published research on zoonotic diseases in Austria, with seven of them involving agents subjected to compulsory surveillance and reporting in humans and/or animals 36 . This outcome underscores an imbalance in research interest, likely influenced by funding opportunities as well as global- and national-level prioritisation, typically based on known incidence and potential impact on human populations. Notably, diseases under European regulatory surveillance, such as those responsible for foodborne outbreaks or those posing a threat to global public health, like the influenza A virus, tend to receive more attention. Such a bias may lead to a skewed assessment of the overall zoonotic risk, especially concerning potentially “neglected” zoonoses such as certain helminth infections (e.g., dirofilariasis, dicrocoeliosis, hepatic capillariasis) 37 . Moreover, research trends show that very few publications in Austria address the environmental compartment, aligning with global observations 38 .
From 1975 to 2022, Austria saw the emergence of eight zoonotic agents, averaging one EID every six years. While there is often an emphasis on viral emergence, particularly considering that RNA viruses pose the most significant threat 39 , our findings offer a different perspective. Within our dataset, six out of eight emerging pathogens in Austria were bacteria and helminths. Notably, two of the emerging bacteria belong to the genus Rickettsia , aligning with the findings of Jones, et al. 4 This highlights the importance of broadening our focus beyond viral threats and acknowledging the substantial role that bacterial and helminthic pathogens play in the landscape of emerging diseases. Moreover, four emerging zoonoses are transmitted by arthropod vectors (WNV, USUV, R. helvetica, R. conorii subsp. raoultii ). As a result of climate change and globalisation, there is a growing likelihood of new arthropod species populations becoming established in Austria, increasing the risk of future EID events 40 . Surprisingly, despite SARS-CoV-2 being notifiable for both humans and animals 41 , 42 , none of the COVID-19-related publications concerning human cases refer to it as a zoonotic disease. Likewise, the sole publication investigating SARS-CoV-2 in Austrian animals did not mention its zoonotic potential 43 .
Within the zoonotic web, multiple zoonotic sources contribute to the maintenance and spread of zoonotic agents. Studying the source-source network of zoonotic agent sharing is necessary to reveal indirect interactions 32 , where one source influences another through shared agents. For example, if an agent is found in two sources, its prevalence in one may affect the other. However, these indirect interactions may lack epidemiological significance if, for instance, immunological or physical barriers prevent agent transfer between sources, such as when the sources do not share similar ecological niches 20 . Besides, many sources found (sero)positive for a zoonotic agent, may not, when taken individually, be able to maintain a sustained persistence of the agent within the network 44 . Nevertheless, as members of a zoonotic source community, interacting with maintenance and non-maintenance sources, they potentially play a role in the zoonotic agent ecology 45 .
We observe that the zoonotic agent sharing network in Austria is organised into six communities. Our results indicate that the community, including humans, the oldest domesticated species (e.g., dog, cat, sheep, cattle, pig 46 ), and synanthropic species (e.g., Norway rat, house mouse) share the most zoonotic agents. This suggests that the highest risk of zoonotic spillover originates from sources within this community. These national-level findings align with results from global studies 2 , 47 . In addition, human-modified environments, such as sandboxes, cluster with humans, domesticated and commensal species, highlighting the role of the shared ecosystem and environmentally persistent stages in the ecology of certain zoonoses 48 . The determinants of the zoonotic source community assembly and composition remain a challenge in disease ecology 5 , 49 . We found evidence that a limited number of highly connected zoonotic agents in the bipartite zoonotic web, such as USUV, S. enterica , WNV, and Influenza A, may, at least partly, drive zoonotic agent sharing community assemblage. The grouping of most food products into one community, predominantly sharing zoonotic agents typically associated with foodborne infections 50 , 51 (21/24 agents, 87.5%, including the five leading causes of foodborne diseases in the EU: Campylobacter, Salmonella, Yersinia , E. coli and Listeria 50 , as well as 10 serovars of Salmonella enterica subsp. enterica) , suggests that anthropogenic activities, particularly those related to food processing and transformation 52 , 53 , may further influence the pattern of assembly within zoonotic source communities. These findings suggest that a combination of local epidemiological, ecological, human-related, and behavioural (e.g., relationship and proximity to human) 2 factors play a key role in shaping zoonotic agent sharing community patterns.
Our findings underscore the presence of central zoonotic sources in the network, demonstrating robust results across three to four centrality metrics after controlling for the research effort. These central zoonotic sources have a higher number of interactions with zoonotic agents, acting as hubs, or bridge different zoonotic source communities in the network, acting as connectors 54 . In particular, some livestock species (e.g., cattle, chicken), companion animals (e.g., dogs, cats, turtles), wildlife (e.g., yellow-necked field mouse, wild boar), and vectors ( Ixodes ) play a crucial role as bridge hosts, through which zoonotic agents can potentially spillover from maintenance (generally wild) host populations or communities to target populations (generally domesticated species or humans) that are usually “protected” through public health or biosecurity measures 25 , 44 , 55 , 56 . Notably, Ixodes ticks are pivotal in the epidemiology and zoonotic spillover of bacteria from the genera Rickettsia, Borrelia , and Babesia . Furthermore, the two communities involving USUV and WNV hosts illustrate the maintenance of zoonotic viruses within partially overlapping host communities. In this subsystem, mosquitoes of the genus Culex play a central role, serving as primary amplification vectors for WNV and USUV within each bird community. In addition, Culex mosquitoes act as bridge vectors between both avian maintenance communities and between these communities and potential mammalian hosts, including humans 57 . These results emphasise the importance of both vector monitoring and testing for pathogens as essential components for the early detection of emerging zoonoses and the establishment of early warning systems.
We present a novel approach based on the identification and quantitative characterisation of specific network structures, named One Health 3-cliques, for estimating the likelihood of zoonotic spillover at human-animal-environment interfaces. This method is flexible and can be applied to any zoonotic web. Our findings demonstrate that there is an increased co-occurrence of zoonotic agents at human-cattle and human-food interfaces, suggesting an elevated likelihood of zoonotic spillover. Notably, human zoonotic infection through consumption of contaminated food is a major public health risk, with Listeria, Salmonella , and Escherichia being the most frequently reported agents in food products across the included publications. Our results further emphasise the critical importance of monitoring zoonotic agents in food-processing environments.
A crucial challenge in formulating One Health surveillance and primary prevention strategies (i.e., at source) 17 for multi-source zoonotic agents, in particular emerging ones, is identifying what is the reservoir of infection 55 , i.e., characterising, within a given context, the “ecologic system in which an infectious agent survives indefinitely” 58 and from which it can be sustainably transmitted to the target population 44 . The goal is to define what could be an optimal (high specificity and sensitivity) sentinel 59 to detect the circulation of a specific zoonotic agent above an acceptable threshold posing a potential transmission risk to the target population (typically human). Identifying sentinels through network metrics should depend on the topology of the network, the infectious agent to be monitored (e.g., endemic versus emerging, transmission route(s)), the (estimated) infection rate, the target population, the objective of the surveillance (e.g., early detection versus prevalence estimation) 60 , 61 , and the specific epidemiological, ecological, and socio-cultural-economic context (e.g., what resources are available, what measures are acceptable, what is the community perception of the disease 62 ). Selecting sentinels that are distant from each other in the network proved to enhance the overall probability of one sentinel being in proximity to an outbreak, thereby increasing the likelihood of detection 63 . For example, distributing the sentinels in different network communities 61 and prioritising surveillance of highly connected nodes in the network 30 (e.g., via regular sampling) would achieve higher performance than randomly selected nodes.
Nodes to be prioritised for surveillance may be different than those used for disease control 63 . Removing central nodes in the network, e.g., via vaccination or culling targeting “bridge” zoonotic sources (i.e., with high betweenness), can significantly reduce the connectivity of the zoonotic web 30 , therefore decreasing the likelihood of zoonotic spillover into the human population. However, betweenness centrality fails to discriminate between zoonotic sources that have high betweenness because they have a lot of connections in the network, such as human and cattle, or sources that really connect two communities, serving as bottlenecks for zoonotic transmission flow 30 (e.g., Ixodes ). Nevertheless, the effectiveness of interventions is intricately connected to the specific system under study and must be tailored to the context. For example, badger culling, equivalent to removing the badger node in the zoonotic web, has shown contrasting results on the prevalence of tuberculosis in cattle in the UK 64 , 65 .
Alternative methodologies have been employed to investigate spillover events. For instance, Grange, et al. 66 ranked the spillover risk from known and newly discovered wildlife-origin viruses using a database of wildlife host-virus associations combined with expert opinion on drivers of spillover. Washburne, et al. 67 utilised percolation models to analyse cross-species transmission, uncovering inherent nonlinearity in spillover rate. In addition, Olival, et al. 8 used a dataset of mammal host-virus associations as a proxy for measuring spillover; using generalised additive models (GAMs), they identified predictors of host viral richness and estimated the number of undiscovered viruses for each mammal species. Missing or unobserved links and nodes frequently occur in collected network data 68 , which can impact the network properties. Diverse methods have been proposed to infer missing links 69 , 70 , 71 and nodes 69 , 72 . Notably, edge prediction accuracy can be enhanced through the use of network community structure 73 . These methods offer valuable mathematical and statistical approaches for future investigations of the zoonotic web, potentially allowing inference of zoonotic agent presence in a source where data has been lacking.
Our study acknowledges several limitations. First, poorly described taxonomic names hinder the precise identification of zoonotic agents or vertebrate hosts at the species level. Likewise, the unspecific description of food origin (e.g., “unspecified” animal), alongside our conservative approach to data validation/cleaning and adherence to authors’ terminology, may have resulted in an inaccurate assessment of the degree centrality for some nodes. For example, Shiga toxin-producing E. coli (STEC) strains could refer to both VTEC and EHEC 74 ; similarly, in the case of a host linked to both Listeria and L. monocytogenes , Listeria could potentially be L. monocytogenes . Imprecise description of the samples and zoonotic agents in publications represents a major limitation to the estimation of the zoonotic risk. Moreover, the single species-single pathogen approach, especially dominant in human medicine 11 , and the tendency to disproportionally investigate zoonotic sources that are closer to humans can result in sample bias and in a skewed distribution of the number of zoonotic agents recorded per source, with human showing the highest number of zoonotic agents, followed by domesticated species. Zoonotic agent detection through environmental sampling remains scarce, potentially limiting result interpretation, particularly in a One Health context. Future zoonotic research could leverage environmental DNA/RNA (eDNA/ eRNA) sequencing for the detection and monitoring of zoonotic agents in the environment 75 , as exemplified by SARS-CoV-2 wastewater-based surveillance 76 . Furthermore, despite efforts to control for research bias, our analysis is inevitably constrained by the existence of zoonotic source-agent associations that are either unknown or not yet published. This constitutes a major challenge in our understanding of zoonotic interactions. Ultimately, broadening the dataset by including additional data on natural infections documented in diverse laboratories (e.g., university laboratories that often investigate a broader range of sources and agents compared to national reference labs) as well as event-based surveillance (EBS) data sourced from various, non-official channels 77 , could significantly enrich the dataset and enhance the depth of the analysis. ProMed-mail reports, for instance, benefit from evaluation by a multidisciplinary team of experts to ensure information reliability and accuracy before publication 78 . Future extensions could also explore conducting targeted searches for each zoonotic agent, based on available global lists 8 , 22 , 23 , 24 , in the literature and international health organisation websites (e.g., World Health Organization, World Organisation for Animal Health). However, the latter may necessitate considerable time and resources; employing automated data extraction methods and tools could improve efficiency 79 , 80 , 81 . In addition, incorporating a temporal dimension to zoonotic source-agent interactions would allow for a more dynamic assessment of the zoonotic transmission chain within and between the communities. This approach could unveil seasonal variations in spillover events 82 as well as mechanisms that link host diversity to disease spread and emergence 83 . Moreover, as data on directionality in transmission is largely unavailable, we used a non-directed network and assumed a symmetrical process in interspecies transmission. This simplification may have limitations in capturing nuances in the dynamics of zoonotic transmission 84 (e.g., WNV can be transmitted from birds to humans via mosquitoes but this transmission process is not reciprocal). Furthermore, our data provides information on infection solely at the species level, overlooking individual variations in shedding, and potentially missing key individuals acting as hubs (“superspreaders”). Finally, controlling for detection method stringency 8 , such as PCR (or other direct detection methods) versus serology, could further refine our findings, allowing us to adjust edge weight within the network.
Here, we show that network analysis represents a cross-disciplinary method for unveiling the intricate web of zoonotic interactions involving multiple sources and infectious agents within an ecological system. In addition to presenting interactions between nodes, a zoonotic web approach enables the identification of influential zoonotic agents and sources that may hold epidemiological significance. Applying this approach across different settings, especially in regions identified as hotspots for zoonotic disease emergence, can expose critical knowledge gaps and reveal how existing epidemiological understanding, shaped by research data availability and funding priorities, may not always reflect on-the-ground realities. Overall, this work emphasises the need for further modelling and empirical studies to explore how maintenance is influenced by multiple source-agent interactions. Establishing efficient and context-adapted One Health network-based surveillance and control strategies requires supplementing the network analysis with multi-source data, ensuring a holistic, multidimensional understanding of the zoonotic web to unravel the complex dynamics of zoonotic transmission chains.
Systematic literature search and data extraction
The systematic literature search was conducted and reported according to the Preferred Reporting Items for Systematic Review and Meta-Analysis (PRISMA) guidelines 85 .
Information about zoonotic agents circulating in Austria is dispersed across scientific papers, reports from the Austrian Agency for Health and Food Safety (AGES), reports from the Federal State Veterinary Services, and student theses. Between 17 July and 23 August 2022, a systematic literature search was conducted using the query (“Zoono*“ AND (“Austria” OR “Österreich”)) in the following databases: PubMed®, Scopus, and vetmed:seeker (internal database of the University of Veterinary Medicine Vienna, Austria), including articles published between the inception of the databases and the date of the search. Furthermore, the publication database of the AGES was searched using the keyword “zoono”. Additional papers found in the reference section of reviews that provided relevant information were also included. Retrieved publications were deduplicated in the reference manager Citavi (Swiss Academic Software. 2023) before the following selection processes.
Titles and abstracts were first screened for relevance using the following inclusion criteria: the publication presented data pertaining to at least one zoonotic disease or agent that was investigated or documented in Austria, and the agent was identified as zoonotic in the paper. Publications were excluded (i) if they did not investigate or describe a zoonotic disease that was identified as such, (ii) if research was not conducted in Austria, (iii) if publications did not describe naturally occurring zoonotic infection, or (iv) if publications described disease physiology or (v) dealt with treatment or methods for pathogen detection. Book chapters, posters, literature reviews, statistical forecasts, and conference proceedings were excluded. Regarding antimicrobial resistant bacteria, papers were included if they specifically explored the animal-human interface and/or the authors referred to zoonotic transmission. To prevent duplication of data, diploma-, master’s-, and doctorate thesis were not included if a peer-reviewed research paper published the same data.
In a second step, the full texts of the previously selected titles/abstracts were screened using the inclusion/exclusion criteria described above. Publications were excluded if they were not in German or English language or did not describe the situation in Austria. When a publication dealt with multiple countries, it was included if it provided specific information on zoonotic diseases in Austria.
The following data was extracted from the selected publications: ( i) Publication data : citation, year of publication, and type of publication; ( ii) Type of study : case study, original research, or national surveillance data; ( iii) Investigated zoonotic agent : agent type (e.g., bacterium, virus, parasite, fungus, prion, or other) and common/scientific names as mentioned in the information source; ( iv) Investigated host : host category, e.g., human, companion animal (defined as domesticated animals possessed by a person for reasons other than food or resource production, including domesticated small rodents or exotic companion animals), livestock (defined as domesticated animals kept for resource and food production), wildlife (defined as free-ranging or captive wild animal species that are not domesticated), common/scientific names as mentioned in the information source, if the zoonotic agent was detected in the host, i.e., seropositive (confirmed by the presence of antibodies), positive (direct detection of the agent), or negative; ( v) Investigated vector: common/scientific names as mentioned in the information source, and if the zoonotic agent was detected in the vector (positive/negative); ( vi) Investigated environmental matrix and if the zoonotic agent was detected in the matrix (positive/negative); ( vii) Investigated food matrix : the specific type of foodstuff investigated, the origin of the food product (animal or plant), and if the zoonotic agent was detected in the foodstuff (positive/negative); ( viii) Epidemiological context: study year, federal state(s), whether the case was imported and most probable origin, whether the zoonotic agent was mentioned as emerging in Austria, and whether specific professional activities were deemed to carry an elevated risk of exposure.
Data curation
First, the data underwent quality control and cleaning procedures where the unique values of each field were checked to search for inaccurate or missing data in the dataset using the R function unique() . Events containing detected errors were manually inspected against the original data source, and, when necessary, the erroneous values were modified, replaced, or removed. Furthermore, for each animal host, vector, and zoonotic agent, common and scientific names, as well as taxonomic classification were resolved against the NCBI Taxonomy database 86 using the R package taxize 87 . If a conflict occurred between the scientific name and/or common name as provided in the information source and the NCBI-resolved name, the information source was cross-referenced and searched for complementary information on the investigated species. When the original source did not provide sufficient details for the identification of a scientific name, the most precise taxonomic denomination was used.
Food categories were generated by combining the food source (e.g., cattle) and the type of food (e.g., meat and meat products). For analytical purposes, foodstuffs designating the same type of food were grouped. For example, “kebab”, “ground meat”, or “rillettes” were coded as “meat and meat product”; “milk” and “milk product” were coded as “dairy”; “egg” and “egg product” were coded as “egg”; “fish”, “fish filet”, or “rollmops” were coded as “marine product”; “salad”, “spices”, “fruit”, or “vegetable” were coded as “plant-based food”. The categories “cheese” (e.g., mozzarella, Brie, Roquefort) and “sausage” (e.g., ham, salami, raw meat sausage) were also added for more accurate representation.
Analysis of the zoonotic web
The dataset was used to create an undirected network representing the web of naturally occurring zoonotic interactions, thereafter called the zoonotic web , depicting the relationships between zoonotic actors. In this network, the zoonotic agents and their zoonotic sources (i.e., vertebrate hosts, arthropod vectors, foodstuffs, and environment) were shown as nodes linked by edges, which represented zoonotic infection (hosts and vectors) or colonisation (food and environmental sources). A link between a zoonotic agent i and a vertebrate host j indicates that agent i was directly or indirectly detected in host j . A link between a zoonotic agent i and a vector j signifies that agent i was identified in vector j , implying that vector j may transmit agent i to a vertebrate host through a bite or mechanically. Likewise, a link between a zoonotic agent i and an environmental matrix j indicates the presence of agent i in environment j , potentially leading to infection of a vertebrate host upon contact. Lastly, a link between a zoonotic agent i and a food matrix j indicates that agent i was detected in food j , which may result in the infection of a vertebrate host through ingestion. In the network, the most specific NCBI-resolved zoonotic agent and host names were employed while arthropod vectors were aggregated at the genus level. The zoonotic web is a bipartite network, i.e., a graph that contains two disjoint sets of nodes, the zoonotic sources and the zoonotic agents, respectively, such that every edge connects the two node sets (i.e., interactions among zoonotic sources or among zoonotic agents were not allowed). The degree centrality (the number of links a node has) was calculated for each node. In the epidemiological context, the node degree centrality for each zoonotic source corresponds to the zoonotic agent richness, i.e., the number of taxa directly or indirectly detected in the zoonotic source. Similarly, the node degree centrality for each zoonotic agent corresponds to the zoonotic source range, i.e., the number of sources from which it has been directly or indirectly evidenced, reflecting its “host” or “zoonotic source” plasticity.
The zoonotic source-agent network was subsequently projected into a one-mode network of zoonotic agent sharing among sources. Edges were weighted by the number of shared zoonotic agents between two sources. By transforming the zoonotic source-agent bipartite network into a source-source unipartite network, a “transmission-potential network“ 29 was created, where sources were linked based on shared zoonotic agents. To account for research biases, we considered, for each source, the total number of zoonotic investigations (i.e., the number of times a source was studied). For instance, if, in one study, a source was investigated annually for three years, we counted three zoonotic investigations. Similarly, if the same source was examined for five zoonotic agents in a single study, we counted it as five zoonotic investigations. This approach provided a more accurate estimation of scientific research effort compared to simply counting the number of studies. We used the number of zoonotic investigations as an estimate of scientific research effort for each source and regressed each edge weight by the Box-Cox transformed number of zoonotic investigations of the least studied source of each edge. The residuals were subsequently rescaled so that the lowest weight value was 1 9 , 10 . After removal of the isolated components in the research effort-adjusted one-mode network of zoonotic agent sharing, we calculated the following node centrality metrics: degree centrality, strength centrality (the sum of the weights of edges to/from a node), weighted betweenness (the number of shortest paths that go through a node, which allows identifying nodes that act as bridges connecting the different communities), and weighted closeness (the average inverse distance to all other nodes) 31 . To calculate the weighted metrics, the edge weight was transformed into cost by dividing 1 by the weight 88 . Node rankings through node centrality metrics were compared using the Kendall correlation test. Average values of the node centrality metrics were also compared between the four zoonotic source categories using the Kruskal-Wallis test. When a difference was evidenced, a pairwise comparison between zoonotic source categories was performed using the Wilcoxon rank sum test; p -values were adjusted following the Benjamini-Hochberg method 89 . Network analyses were performed using the R packages igraph 90 and bipartite 91 .
Community detection
We used the Leiden algorithm 92 , which relies on a measure called modularity 93 , to detect communities of zoonotic agent sharing within the research-adjusted one-mode network of zoonotic sources. The method aims to optimise modularity by maximising the difference between the actual and expected number of edges within communities. The Leiden algorithm is considered as an improvement over the Louvain algorithm 94 . It comprises three distinct steps: initial optimisation of modularity, subsequent refinement of the partition, and a third step focusing on the community aggregation process 92 . Notably, by refining the local partition in each community, the Leiden algorithm demonstrates enhanced stability in community detection and offers more efficient computation time compared to the Louvain algorithm 94 .
Exploring network One Health cliques
We investigated the circulation of zoonotic agents at human-animal-environment interfaces within the research effort-adjusted network of zoonotic agent sharing by searching “One Health” 3-cliques in the network structure. A clique is a fully connected subgraph within the network. A 3-clique is a set of three nodes all pair-wisely connected to each other, therefore forming a triangle 31 . We were interested in 3-cliques that included nodes representing the three traditional One Health compartments, i.e., animal (173 nodes), human (one node), and environment (15 nodes). Plant-based foodstuffs, invertebrate vectors, and any environmental matrices (including surfaces and tools in food processing plants) were included into the compartment “environment” while food products of animal origin were considered within the “animal” compartment. We ranked the One Health cliques by their total edge weight, i.e., the sum of the edge weights between all pairs of nodes (zoonotic sources) within the clique, corresponding to the research effort-adjusted sum of zoonotic agents shared between all pairs of One Health compartments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw dataset generated in this study, as well as its cleaned and validated version, are available in the Supplementary Code. The data has also been archived in the study repository on figshare with the identifier: https://doi.org/10.6084/m9.figshare.25306177 95 .
Code availability
All analyses were conducted in R Statistical Software version 4.3.0 (2023-04-21) “Already Tomorrow”. The documented R scripts used for data cleaning, validation, processing, and analysis, are available in Supplementary Code. The READ.ME file contains the necessary instructions to run the code and replicate our results. These files have also been archived in the study repository on figshare 95 .
Kock, R. & Caceres-Escobar, H. Situation analysis on the roles and risks of wildlife in the emergence of human infectious diseases. 112 (IUCN Species Survival Commission (SSC), Gland, Switzerland, 2022).
Morand, S., McIntyre, K. M. & Baylis, M. Domesticated animals and human infectious diseases of zoonotic origins: domestication time matters. Infect. Genet. Evol. 24 , 76–81 (2014).
Article PubMed Google Scholar
McNeill, W. Plagues and peoples . (Anchor Book Editions, 1976).
Jones, K. E. et al. Global trends in emerging infectious diseases. Nature 451 , 990–993 (2008).
Article ADS CAS PubMed PubMed Central Google Scholar
Buhnerkempe, M. G. et al. Eight challenges in modelling disease ecology in multi-host, multi-agent systems. Epidemics 10 , 26–30 (2015).
Hassell, J. M., Begon, M., Ward, M. J. & Fèvre, E. M. Urbanization and disease emergence: dynamics at the wildlife-livestock-human interface. Trends Ecol. Evol. 32 , 55–67 (2017).
Article PubMed PubMed Central Google Scholar
Dobson, A. Population dynamics of pathogens with multiple host species. Am. Nat. 164 , S64–S78 (2004).
Olival, K. J. et al. Host and viral traits predict zoonotic spillover from mammals. Nature 546 , 646–650 (2017).
Gómez, J. M., Nunn, C. L. & Verdú, M. Centrality in primate–parasite networks reveals the potential for the transmission of emerging infectious diseases to humans. PNAS 110 , 7738–7741 (2013).
Article ADS PubMed PubMed Central Google Scholar
Luis, A. D. et al. Network analysis of host–virus communities in bats and rodents reveals determinants of cross-species transmission. Ecol. Lett. 18 , 1153–1162 (2015).
Cleaveland, S., Laurenson, M. K. & Taylor, L. H. Diseases of humans and their domestic mammals: pathogen characteristics, host range and the risk of emergence. Philos. Trans. R. Soc. Lond. B Biol. Sci. 356 , 991–999 (2001).
Article CAS PubMed PubMed Central Google Scholar
Ghai, R. R. et al. A generalizable One Health framework for the control of zoonotic diseases. Sci. Rep. 12 , 8588 (2022).
Karesh, W. B. et al. Ecology of zoonoses: natural and unnatural histories. Lancet 380 , 1936–1945 (2012).
Hirst, K. M. & Halsey, S. J. Bacterial zoonoses impacts to conservation of wildlife populations: a global synthesis. Front. Conserv. Sci . 4 , https://doi.org/10.3389/fcosc.2023.1218153 (2023).
Bernstein, A. S. et al. The costs and benefits of primary prevention of zoonotic pandemics. Sci. Adv. 8 , eabl4183 (2022).
British Academy. The COVID Decade: understanding the long-term societal impacts of COVID-19. 172 (The British Academy, London, United Kingdom, 2021).
Markotter, W. et al. Prevention of zoonotic spillover: From relying on response to reducing the risk at source. PLoS Pathog. 19 , e1011504 (2023).
LoGiudice, K., Ostfeld, R. S., Schmidt, K. A. & Keesing, F. The ecology of infectious disease: effects of host diversity and community composition on Lyme disease risk. Proc. Natl. Acad. Sci. USA 100 , 567–571 (2003).
Keesing, F. et al. Impacts of biodiversity on the emergence and transmission of infectious diseases. Nature 468 , 647–652 (2010).
Plowright, R. K. et al. Pathways to zoonotic spillover. Nat. Rev. Microbiol. 15 , 502–510 (2017).
Lloyd-Smith, J. O. et al. Epidemic dynamics at the human-animal interface. Science 326 , 1362–1367 (2009).
Mollentze, N. & Streicker, D. G. Viral zoonotic risk is homogenous among taxonomic orders of mammalian and avian reservoir hosts. Proc. Natl. Acad. Sci. USA 117 , 9423–9430 (2020).
Carlson, C. J. et al. The global virome in one network (VIRION): an atlas of vertebrate-virus associations. mBio 13 , e02985–02921 (2022).
Wardeh, M., Risley, C., McIntyre, M. K., Setzkorn, C. & Baylis, M. Database of host-pathogen and related species interactions, and their global distribution. Sci. Data 2 , 150049 (2015).
Jones, B. A. et al. Zoonosis emergence linked to agricultural intensification and environmental change. PNAS 110 , 8399–8404 (2013).
Geiser, E. Wie viele tierarten leben in Österreich? Erfassung, Hochrechnung und Abschätzung. Verh. Zool. Bot. Ges. Wien. 135 , 81–93 (1998).
Google Scholar
Statistik Austria. https://www.statistik.at/en (2023).
Bascompte, J. Disentangling the web of life. Science 325 , 416–419 (2009).
Article ADS MathSciNet CAS PubMed Google Scholar
Pilosof, S., Morand, S., Krasnov, B. R. & Nunn, C. L. Potential parasite transmission in multi-host networks based on parasite sharing. PLoS ONE 10 , e0117909 (2015).
VanderWaal, K. L., Atwill, E. R., Isbell, L. A. & McCowan, B. Quantifying microbe transmission networks for wild and domestic ungulates in Kenya. Biol. Conserv. 169 , 136–146 (2014).
Article Google Scholar
Newman, M. Networks: An Introduction 2nd edn, (Oxford University Press, Inc., 2018).
Simmons, B. I. et al. Motifs in bipartite ecological networks: uncovering indirect interactions. Oikos 128 , 154–170 (2019).
Article ADS Google Scholar
Delmas, E. et al. Analysing ecological networks of species interactions. Biol. Rev. 94 , 16–36 (2019).
Granovetter, M. S. The strength of weak ties. Am. J. Sociol. 78 , 1360–1380 (1973).
Combes, C. Parasitism: the ecology and evolution of intimate interactions . (University of Chicago Press, 2001).
Bundeskanzleramt Österreich. Bundesgesetz zur Überwachung von Zoonosen und Zoonoseerregern (Zoonosengesetz). https://www.ris.bka.gv.at/eli/bgbl/I/2005/128 (2005).
Casulli, A. et al. Unveiling the incidences and trends of the neglected zoonosis cystic echinococcosis in Europe: a systematic review from the MEmE project. Lancet Infect. Dis. 23 , e95–e107 (2023).
Essack, S. Y. Environment: the neglected component of the One Health triad. Lancet Planet. Health 2 , e238–e239 (2018).
Keusch, G. T. et al. Pandemic origins and a One Health approach to preparedness and prevention: Solutions based on SARS-CoV-2 and other RNA viruses. PNAS 119 , e2202871119 (2022).
Semenza, J. C. & Suk, J. E. Vector-borne diseases and climate change: a European perspective. FEMS Microbiol. Lett. 365 , https://doi.org/10.1093/femsle/fnx244 (2018).
Bundeskanzleramt Österreich. Epidemiegesetz (BGBl. Nr. 186/1950). https://www.ris.bka.gv.at/Dokumente/BgblPdf/1950_186_0/1950_186_0.pdf (1950).
World Organisation for Animal Health. Terrestrial animal health code. https://www.woah.org/en/what-we-do/standards/codes-and-manuals/terrestrial-code-online-access/ (2022).
Moreira-Soto, A. et al. Serological evidence that SARS-CoV-2 has not emerged in deer in Germany or Austria during the COVID-19 pandemic. Microorganisms 10 , https://doi.org/10.3390/microorganisms10040748 (2022).
Roberts, M. G. & Heesterbeek, J. A. P. Characterizing reservoirs of infection and the maintenance of pathogens in ecosystems. J. R. Soc. Interface 17 , 20190540 (2020).
Bordes, F., Caron, A., Blasdell, K., de Garine-Wichatitsky, M. & Morand, S. Forecasting potential emergence of zoonotic diseases in South-East Asia: network analysis identifies key rodent hosts. J. Appl. Ecol. 54 , 691–700 (2017).
McHugo, G. P., Dover, M. J. & MacHugh, D. E. Unlocking the origins and biology of domestic animals using ancient DNA and paleogenomics. BMC Biol. 17 , 98 (2019).
Wells, K., Morand, S., Wardeh, M. & Baylis, M. Distinct spread of DNA and RNA viruses among mammals amid prominent role of domestic species. Glob. Ecol. Biogeogr. 29 , 470–481 (2020).
Hopkins, S. R. et al. Environmental persistence of the world’s most burdensome infectious and parasitic diseases. Front. Public Health 10 , https://doi.org/10.3389/fpubh.2022.892366 (2022).
Johnson, P. T. J., de Roode, J. C. & Fenton, A. Why infectious disease research needs community ecology. Science 349 , 1259504 (2015).
European Food Safety Authority & European Centre for Disease Prevention. The European Union One Health 2022 Zoonoses Report. EFSA J. 21 , p211202 (2023).
Hald, T. et al. World Health Organization estimates of the relative contributions of food to the burden of disease due to selected foodborne hazards: a structured expert elicitation. PLoS ONE 11 , e0145839 (2016).
Sun, L. et al. Epidemiology of foodborne disease outbreaks caused by nontyphoidal Salmonella in Zhejiang Province, China, 2010-2019. Foodborne Pathog. Dis. 18 , 880–886 (2021).
Article CAS PubMed Google Scholar
Gallo, M., Ferrara, L., Calogero, A., Montesano, D. & Naviglio, D. Relationships between food and diseases: What to know to ensure food safety. Food Res. Int. 137 , 109414 (2020).
Mello, M. A. R., Bezerra, E. L. S. & Machado, I. C. Functional roles of Centridini oil bees and Malpighiaceae oil flowers in biome-wide pollination networks. Biotropica 45 , 45–53 (2013).
Haydon, D. T., Cleaveland, S., Taylor, L. H. & Laurenson, M. K. Identifying reservoirs of infection: a conceptual and practical challenge. Emerg. Infect. Dis. 8 , 1468–1473 (2002).
Caron, A., Cappelle, J., Cumming, G. S., de Garine-Wichatitsky, M. & Gaidet, N. Bridge hosts, a missing link for disease ecology in multi-host systems. Vet. Res. 46 , 83 (2015).
Kilpatrick, A. M. et al. West Nile virus risk assessment and the bridge vector paradigm. Emerg. Infect. Dis. 11 , 425–429 (2005).
Ashford, R. W. When is a reservoir not a reservoir? Emerg. Infect. Dis. 9 , 1495–1496 (2003).
Halliday, J. E. et al. A framework for evaluating animals as sentinels for infectious disease surveillance. J. R. Soc. Interface 4 , 973–984 (2007).
Radici, A., Bevacqua, D., Miele, L. & Martinetti, D. Network-thinking to optimize surveillance and control of crop parasites. A review. Preprint at https://doi.org/10.48550/arXiv.42310.07442 (2023).
Colman, E., Holme, P., Sayama, H. & Gershenson, C. Efficient sentinel surveillance strategies for preventing epidemics on networks. PLoS Comput. Biol. 15 , e1007517 (2019).
Guenin, M.-J. et al. A participatory epidemiological and One Health approach to explore the community’s capacity to detect emerging zoonoses and surveillance network opportunities in the forest region of Guinea. PLoS Negl. Trop. Dis. 16 , e0010462 (2022).
Holme, P. Three faces of node importance in network epidemiology: Exact results for small graphs. Phys. Rev. E 96 , 062305 (2017).
Griffin, J. M. et al. The impact of badger removal on the control of tuberculosis in cattle herds in Ireland. Prev. Vet. Med. 67 , 237–266 (2005).
Article ADS CAS PubMed Google Scholar
Donnelly, C. A. et al. Impact of localized badger culling on tuberculosis incidence in British cattle. Nature 426 , 834–837 (2003).
Grange, Z. L. et al. Ranking the risk of animal-to-human spillover for newly discovered viruses. PNAS 118 , e2002324118 (2021).
Washburne, A. D. et al. Percolation models of pathogen spillover. Philos. Trans. R. Soc. B Biol. Sci. 374 , 20180331 (2019).
Lü, L. & Zhou, T. Link prediction in complex networks: a survey. Phys. A 390 , 1150–1170 (2011).
Kim, M. & Leskovec, J. In Proceedings of the 2011 SIAM International Conference on Data Mining (SDM). 47–58 (2011).
Clauset, A., Moore, C. & Newman, M. E. J. Hierarchical structure and the prediction of missing links in networks. Nature 453 , 98–101 (2008).
Guimerà, R. & Sales-Pardo, M. Missing and spurious interactions and the reconstruction of complex networks. PNAS 106 , 22073–22078 (2009).
Liu, C., Li, Z. & Zhou, L. Missing nodes detection for complex networks based on graph convolutional networks. J. Ambient Intell. Humaniz. Comput. 14 , 9145–9158 (2023).
Yan, B. & Gregory, S. Finding missing edges in networks based on their community structure. Phys. Rev. E 85 , 056112 (2012).
Kaper, J. B., Nataro, J. P. & Mobley, H. L. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2 , 123–140 (2004).
Bass, D., Christison, K. W., Stentiford, G. D., Cook, L. S. J. & Hartikainen, H. Environmental DNA/RNA for pathogen and parasite detection, surveillance, and ecology. Trends Parasitol. 39 , 285–304 (2023).
Amman, F. et al. Viral variant-resolved wastewater surveillance of SARS-CoV-2 at national scale. Nat. Biotechnol. 40 , 1814–1822 (2022).
Dub, T. et al. Epidemic intelligence activities among national public and animal health agencies: a European cross-sectional study. BMC Public Health 23 , 1488 (2023).
Yu, V. L. & Madoff, L. C. ProMED-mail: An early warning system for emerging diseases. Clin. Infect. Dis. 39 , 227–232 (2004).
Gutiérrez-Sacristán, A. et al. Text mining and expert curation to develop a database on psychiatric diseases and their genes. Database 2017 , bax043 (2017).
Paraskevopoulos, S., Smeets, P., Tian, X. & Medema, G. Using Artificial Intelligence to extract information on pathogen characteristics from scientific publications. Int. J. Hyg. Environ. Health 245 , 114018 (2022).
Walker, V. R. et al. Evaluation of a semi-automated data extraction tool for public health literature-based reviews: Dextr. Environ. Int. 159 , 107025 (2022).
Nandi, A. & Allen, L. J. S. Probability of a zoonotic spillover with seasonal variation. Infect. Dis. Model 6 , 514–531 (2021).
PubMed PubMed Central Google Scholar
Halliday, F. W., Heckman, R. W., Wilfahrt, P. A. & Mitchell, C. E. Past is prologue: host community assembly and the risk of infectious disease over time. Ecol. Lett. 22 , 138–148 (2019).
Fagre, A. C. et al. Assessing the risk of human-to-wildlife pathogen transmission for conservation and public health. Ecol. Lett. 25 , 1534–1549 (2022).
Page, M. J. et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. Syst. Rev. 10 , 89 (2021).
Schoch, C. L. et al. NCBI Taxonomy: a comprehensive update on curation, resources and tools. Database 2020 , baaa062 (2020).
Chamberlain, S. A. & Szöcs, E. taxize: taxonomic search and retrieval in R. F1000Res 2 , 191 (2013).
Newman, M. E. J. Scientific collaboration networks. II. Shortest paths, weighted networks, and centrality. Phys. Rev. E 64 , 016132 (2001).
Article ADS CAS Google Scholar
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 57 , 289–300 (1995).
Article MathSciNet Google Scholar
Csardi, G. & Nepusz, T. The igraph software package for complex network research. Int. J. Complex Syst . 1695 , 1–9 (2006).
Dormann, C., Gruber, B. & Fründ, J. Introducing the bipartite package: Analysing ecological networks. R News 8 , 11–9 (2008).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9 , 5233 (2019).
Newman, M. E. J. & Girvan, M. Finding and evaluating community structure in networks. Phys. Rev. E 69 , 026113 (2004).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. Theory Exp. 2008 , P10008 (2008).
Desvars-Larrive, A. et al. Repository for the paper: A One Health framework for exploring zoonotic interactions demonstrated through a case study. Figshare https://doi.org/10.6084/m9.figshare.25306177.v1 (2024).
Download references
Acknowledgements
The authors thank Sina Sajjadi and Rafael Prieto-Curiel for their valuable insights and advice on network analysis.
Author information
Authors and affiliations.
Centre for Food Science and Veterinary Public Health, Clinical Department for Farm Animals and Food System Science, University of Veterinary Medicine Vienna, Vienna, Austria
Amélie Desvars-Larrive, Anna Elisabeth Vogl, Gavrila Amadea Puspitarani & Annemarie Käsbohrer
Complexity Science Hub, Vienna, Austria
Amélie Desvars-Larrive, Gavrila Amadea Puspitarani & Liuhuaying Yang
Centre of Pathobiology, Department of Biological Sciences and Pathobiology, University of Veterinary Medicine Vienna, Vienna, Austria
Anja Joachim
You can also search for this author in PubMed Google Scholar
Contributions
Initial concept: A.D-L.; Analysis design: A.D-L.; Data collection: A.V. and A.D-L.; Data collation and processing: A.D-L. and A.V.; Data analysis and modelling: A.D-L. and G.A.P.; Visualisation: A.D-L., L.Y. and G.A.P.; Writing (initial draft): A.D-L. and A.V.; Writing (review and editing): A.D-L., A.V., G.A.P., L.Y., A.J. and A.K.
Corresponding author
Correspondence to Amélie Desvars-Larrive .
Ethics declarations
Competing interests.
The authors declare no competing interests.
Peer review
Peer review information.
Nature Communications thanks Younjung Kim, Thierry Lefrançois and Marisa Peyre for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information, peer review file, description of additional supplementary files, supplementary code, reporting summary, rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/ .
Reprints and permissions
About this article
Cite this article.
Desvars-Larrive, A., Vogl, A.E., Puspitarani, G.A. et al. A One Health framework for exploring zoonotic interactions demonstrated through a case study. Nat Commun 15 , 5650 (2024). https://doi.org/10.1038/s41467-024-49967-7
Download citation
Received : 26 February 2024
Accepted : 24 June 2024
Published : 15 July 2024
DOI : https://doi.org/10.1038/s41467-024-49967-7
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
By submitting a comment you agree to abide by our Terms and Community Guidelines . If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.

IMAGES
VIDEO
COMMENTS
Case Study Questions for Class 10 Maths Chapter 6 - Triangles. CASE STUDY 1: Vijay is trying to find the average height of a tower near his house. He is using the properties of similar triangles ...
CBSE 10th Standard Maths Subject Triangles Case Study Questions With Solution 2021. An aeroplane leaves an airport and flies due north at a speed of 1200km /hr. At the same time, another aeroplane leaves the same station and flies due west at the speed of 1500 km/hr as shown below. After \ (1 \frac {1} {2}\) hr both the aeroplanes reaches at ...
Students looking for Case Study on Triangles Class 10 Maths can use this page to download the PDF file. The case study questions on Triangles are based on the CBSE Class 10 Maths Syllabus, and therefore, referring to the Triangles case study questions enable students to gain the appropriate knowledge and prepare better for the Class 10 Maths ...
Triangles Case Study Questions With answers. Here, we have provided case-based/passage-based questions for Class 10 Maths Chapter 6 Triangles. Case Study/Passage-Based Questions. Question 1: Meenal was trying to find the height of the tower near his house. She is using the properties of similar triangles. The height of Meenal's house is 20 m.
These Case Study and Passage Based questions are published by the experts of CBSE Experts for the students of CBSE Class 10 so that they can score 100% on Boards. CBSE Class 10 Mathematics Exam 2024 will have a set of questions based on case studies in the form of MCQs. The CBSE Class 10 Mathematics Question Bank on Case Studies, provided in ...
Question 5 - Case Based Questions (MCQ) - Chapter 6 Class 10 Triangles Last updated at April 16, 2024 by ... none of these Since this question involves Pythagoras Theorem Which is in Triangles Class 10 So, the correct answer is (a) Show More. Next: NCERT Exemplar - MCQ → Go Ad-free Chapter 6 Class 10 Triangles; Serial order wise ...
By QB365 on 09 Sep, 2022 . QB365 provides a detailed and simple solution for every Possible Case Study Questions in Class 10th Maths Subject - Triangles, CBSE. It will help Students to get more practice questions, Students can Practice these question papers in addition to score best marks.
Updated for new NCERT 2024-2025 Edition. Get NCERT Solutions of Chapter 6 Class 10 Triangles free at teachoo. Solutions to all NCERT Exercise Questions, Examples, Theorems, Optional Exercises are available with Videos of each and every question. We have studied Congruency of Triangles in Class 9.
Triangles Case Study Questions (CSQ's) Practice Tests. Timed Tests. Select the number of questions for the test: Select the number of questions for the test: TopperLearning provides a complete collection of case studies for CBSE Class 10 Maths Triangles chapter. Improve your understanding of biological concepts and develop problem-solving ...
NCERT Solutions Class 10 Maths Chapter 6, Triangles, is part of the Unit Geometry, which constitutes 15 marks of the total marks of 80. On the basis of the updated CBSE Class 10 Syllabus for 2023-24, this chapter belongs to the Unit-Geometry and has the second-highest weightage. Hence, having a clear understanding of the concepts, theorems and ...
Triangles Case Study Questions With Answers. Here, we have provided case-based/passage-based questions for Class 10 Maths Chapter 6 Triangles. Case Study/Passage-Based Questions. Case Study 1: Meenal was trying to find the height of the tower near his house. She is using the properties of similar triangles. The height of Meenal's house is 20 m.
The Case Based Questions: Triangles is an invaluable resource that delves deep into the core of the Class 10 exam. These study notes are curated by experts and cover all the essential topics and concepts, making your preparation more efficient and effective.
This video explains the detailed solution and explanation of Case Study Based Questions related to Chapter 6 Triangles which include the concepts of Similar ... CBSE Exam, class 10
4. (Motivate) If the corresponding sides of two triangles are proportional, their corresponding angles are equal and the two triangles are similar. 5. (Motivate) If one angle of a triangle is equal to one angle of another triangle and the sides including these angles are proportional, the two triangles are similar. 1.
Join Now! Case Study Questions for Class 10 Maths Chapter 6 Triangles. Case Study Questions: Question 1: Vijay is trying to find the average height of a tower near his house. He is using the properties of similar triangles. The height of Vijay's house if 20 m when Vijay's house casts a shadow 10 m long on the ground.
CBSE Board has introduced the case study questions for the ongoing academic session 2021-22. The board will ask the paper on the basis of a different exam pattern which has been introduced this year where 50% syllabus is occupied for MCQ for Term 1 exam. Selfstudys has provided below the chapter-wise questions for CBSE Class 10 Maths.
NCERT Exemplar Class 10 Maths Chapter 6 Exercise - 6.3. Question 1. In a ∆PQR, PR 2 - PQ 2 = QR 2 and M is a point on side PR such that QM⊥ PR. Prove that QM 2 = PM × MR. Solution: Given, In ∆PQR, PR 2 - PQ 2 = QR 2 and QM ⊥ PR.
Ex 6.4 Class 10 Maths Question 8. Tick the correct answer and justify (i) ABC and BDE are two equilateral triangles such that D is the mid-point of BC. Ratio of the areas of triangles ABC and BDE is (a) 2 :1 (b) 1:2 (c) 4 :1 (d) 1:4 (ii) Sides of two similar triangles are in the ratio 4 : 9.
Triangles Class 10 Maths Important Questions Short Answer-I (2 Marks) 12. In an equilateral triangle of side 3√3 cm find the length of the altitude. Solution. Let ∆ABC be an equilateral triangle of side 3√3 cm and AD is altitude which is also a perpendicular bisector of side BC. This is shown in figure given below. 13.
Help them to answer these questions. (i) If an insect (small ant) walks 24 m from H to F, then walks 6 m to reach at M, then walks 4 m to reach at L and finally crossing K, reached at J. Find the distance between initial and final position of insect. (a) 25m. (b) 26m.
MCQ Questions for Classs 10 Maths Chapter 6 Triangle. There are total 6 exercises in the chapter in which last one is optional. NCERT Solutions for Class 10 Maths Chapter 6 Triangles is also available exercisewise that is given below. Exercise 6.1. Exercise 6.2.
NCERT Solutions Class 10 Maths Chapter 6 Triangles are provided here to help the students of CBSE class 10. Our expert teachers prepared all these solutions as per the latest CBSE syllabus and guidelines. In this chapter, we have discussed the similarity of triangles, criterion for similarity of triangles, areas of similar triangles and ...
Note: Check more Case Based Questions for - Triangles Class 10 Note: Check ... Question 18 Case Study Based- 2 SCALE FACTOR AND SIMILARITY SCALE FACTOR A scale drawing of an object is the same shape as the object but a different size. The scale of a drawing is a comparison of the length used on a drawing to the length it represents.
Zoonotic disease systems comprise complex dynamics between multiple animal hosts, vectors, environmental and food sources, and humans. In this study, the authors characterise this 'zoonotic web ...
Innovative teaching strategies using questioning for experimental teaching of genetics: a case study. Fang Ma Key Laboratory of Resource Utilization of Agricultural Solid Waste in Gansu Province, Tianshui Normal University, ... The pre-class preview link is set up, the preview content is arranged in advance, and the preview task is clear, which ...